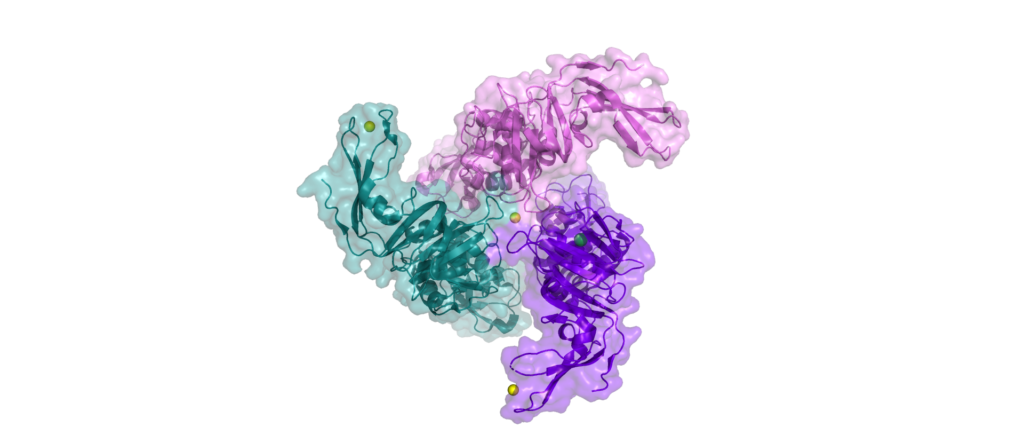
Nsp3 is made up of over 2000 amino acid residues.
As the largest protein encoded by the coronavirus’s genome, untangling its structure and function poses a huge task.
No complete structure of nsp3 has been solved, only single domains, of which nsp13 possesses 10 or more (depending on how you count). It has several transmembrane domains, making it a membrane-bound protein. It also interacts with the immune system in various ways, suppressing the cell’s immune response.
The most studied domain acts tocleave polypeptides, just like the main protease.
When the RNA of SARS-CoV-2 is translated into proteins for the first time, just after viral entry, a long protein chain is produced, a poly-peptide, which needs to be cleaved into functional parts. The first three functional parts, nsp1, nsp2, and nsp3 are cleaved by the papain-like protease domain of nsp3.
Without this activity, the virus cannot replicate and the infection cannot take hold. Hence, this domain is an important drug target.