The instructions and files below will allow you to create your own model of the virus! All you need is some spare time and a 3D printer. In addition, those without access to a 3D printer can still use the STL files to request printing from external services and then follow the instructions on painting and assembling the same way. We do hope that this model will make the virus more tangible, and that the model will not only be printed as a private project, but also be used for outreach activities and in educational institutions.
Our design is based on the best scientific evidence available. Not only are the shapes of the various proteins as true as we can make them, but their numbers as well as the overall size of the virion match experimental results on a scale of 1:1,000,000. If you want to know more about it, please look here. Once you have built a model from our design you will have a good representation of what one of these virions is expected to look like, after being scaled up by a factor of 1,000,000. Therefore 1 mm on the model represents 1 nm (10 Å). (By the way, this would make the RNA that is inside the virus hull 10 metres long and 1 mm thick, and the nucleocapsid around which the RNA is coiled would be about 1 metre and 1 cm in diameter.
We have also designed a scale model of the human anti-body that binds to the spike protein. This is available alongside the virus model and can be attached to the spike protein as desired. For easier printing, painting, and assembly, the virus structure has been broken down into 4 unique components:

To date the structures have been printed successfully on several Fused Deposition Modelling (FDM) printers (Rostok MAX v2 & Prusa I3 MK3 printers), and we anticipate the even higher quality structures will be feasible with alternate methods, such as stereolithography (watch this space). Let us know in the comments! Each of the parts is available in STL format and should be printable through any suitable slicer software. Personal discretion is advised when setting up the prints, as the exact details may differ depending on conditions and equipment. The procedure outlined below will serve as a good starting point.
Printing of the component parts
The first step is to print the individual components. For the virion parts this is very straight forward as the flat surface negates the need for supports. The virion objects can be printed with the minimum infill for support, though infill of 10% is recommended for rigidity.
The other parts (spike proteins and antibodies) provide a more challenging print. The spike protein must be printed 95 times to complete the model, and users can arrange these individually, or using 4 prints of 25x STL file. It is recommended that the spike protein is printed with the crown facing towards the print bed to maximize the support between the bed and eliminating the need to remove supports from the thin delicate stem.
A dual extruder printer would be ideal for spike printing as it would allow supports to be printed in a water soluble plastic, speeding up post-processing. In either case, printing individual or at least fewer spikes with greater spacing generally produces nicer objects that are easier to work with at the price of longer printing time. Indeed, there is a general trade-off between the convenience of the print set-up and the amount of post-processing and tidying needed for all 3D printing tasks, and one must find a compromise which satisfies them.
As stated above, we used FDM printing and ubiquitous poly-lactic acid (PLA), which made the post-processing easier.
Post-processing
Regardless of the approach taken for printing, some amount of tidying will typically be needed to get the objects ready for assembly. Removing the supports can be done with a pair of plyers, while the smaller artifacts and issues will need brushing off or sanding. A dental pick can be quite useful.
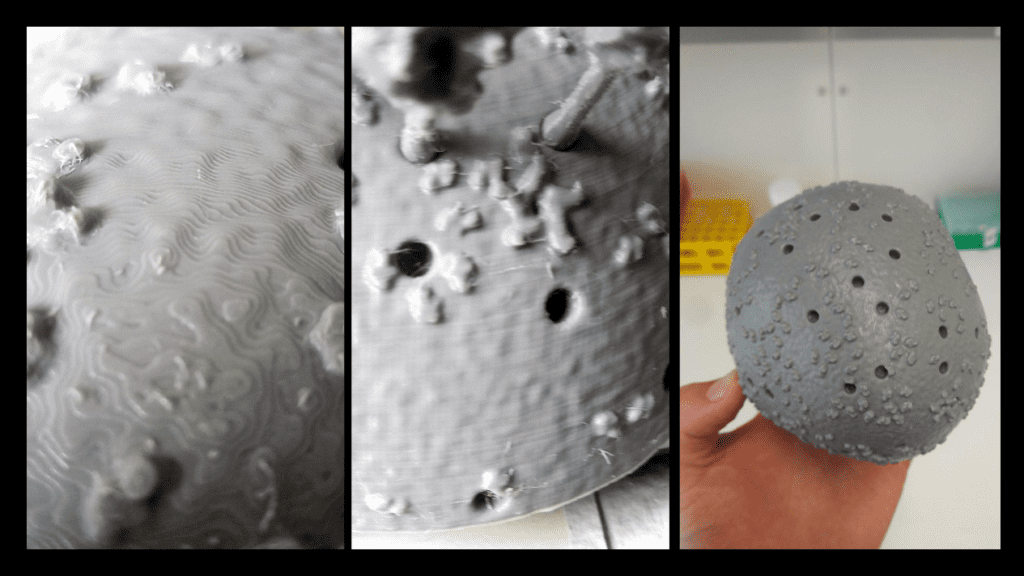

For PLA, we found the best thing to clean and smooth the surfaces (after support removal), is ethyl acetate. Ethyl acetate dissolves the plastic, breaking down the small extrusion artifacts on the surfaces. This can be used in many ways. We found it best to leave the parts in a sealed ethyl acetate vapour environment, such as a stainless steel pot, which should be cleaned carefully afterwards. This technique results in the most even and clean results, though will take up to a few days to fully smooth each object. The faster method, is to simply submerge the small objects in ethyl acetate for 10-30 seconds, and then remove each object, leaving them to dry out on a surface. For the larger virion parts, the surface can be smoothed by rubbing it down with a cloth damped with ethyl acetate. Ethyl acetate was also used to “weld” the two virion parts together. A small amount was dropped onto the flat surfaces on each section, before the two were pressed together until the plastic fused to become a single object. The seam was then smoothed down using the same process as before. Where one cannot get ethyl acetate from a lab or pharmacy, acetone-free nail-polish remover offers a commercially accessible alternative. you should be using safety glasses and suitable (!) gloves when handling ethyl acetate, ventilate the room well and if there was skin contact use a skin cream after hand washing.
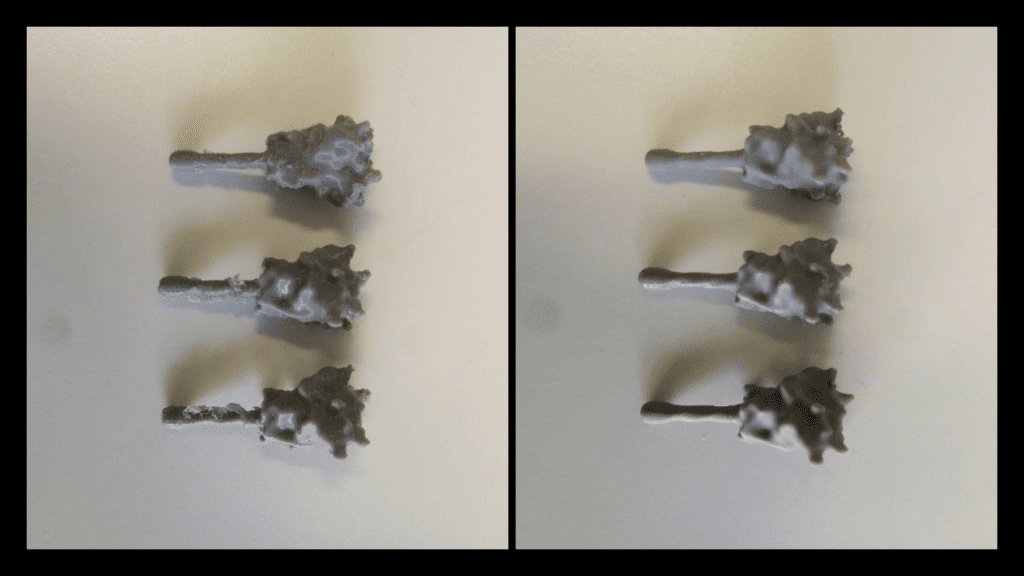

It is worth noting that for the other common 3D printing material acrylonitrile butadiene styrene (ABS) or acetone may produce the same results.
Painting and Gluing
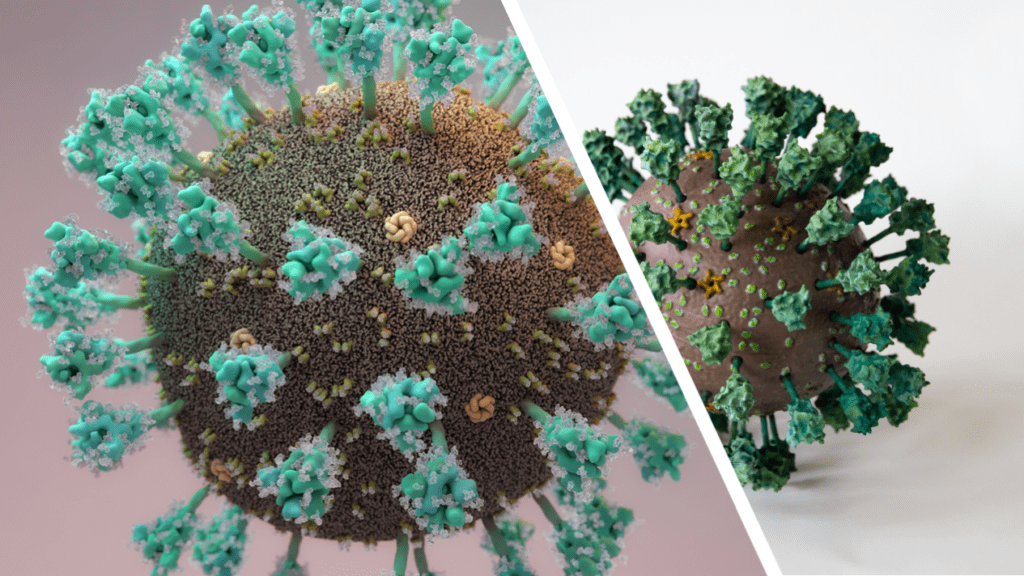

As with printing, painting methods and colours is down to personal preference, and here we outline our attempt, which was guided by the illustration by Thomas Splettstoesser as close as possible (see Fig. 3).
The parts were first treated with a primer to help the paint stick to the model. This also acts as a nice even Basecoat. When working with either primer or, as discussed later, an airbrush, one should consider safety: try to do as much as you can in a ventilated space, wearing safety goggles, gloves and a mask. Paint spraying produces a great number of fine particles which you don`t want to breathe in.
For us, the painting process was performed largely with an airbrush, and we highly recommend using one where available, due to the amount of painting required and surface complexity. Where not available, it can of course be done with just a simple brush which will take more time and a higher skill level.
All layer colours, medium thinner, base colours, primer and varnish we used were from Citadel painting. Here is an outline of the specific Citadel colours and materials we used for the model in the figures:
- Lime: “Moot green”
- Yellow: “Yriel Yellow”
- Grey: “Dawnstone”
- Wheat: “Baneblade Brown”
- Chocolate: “Doombull Brown”
- Aqua: “Gauss Blaster Green”
- Teal: “Kabalite Green”
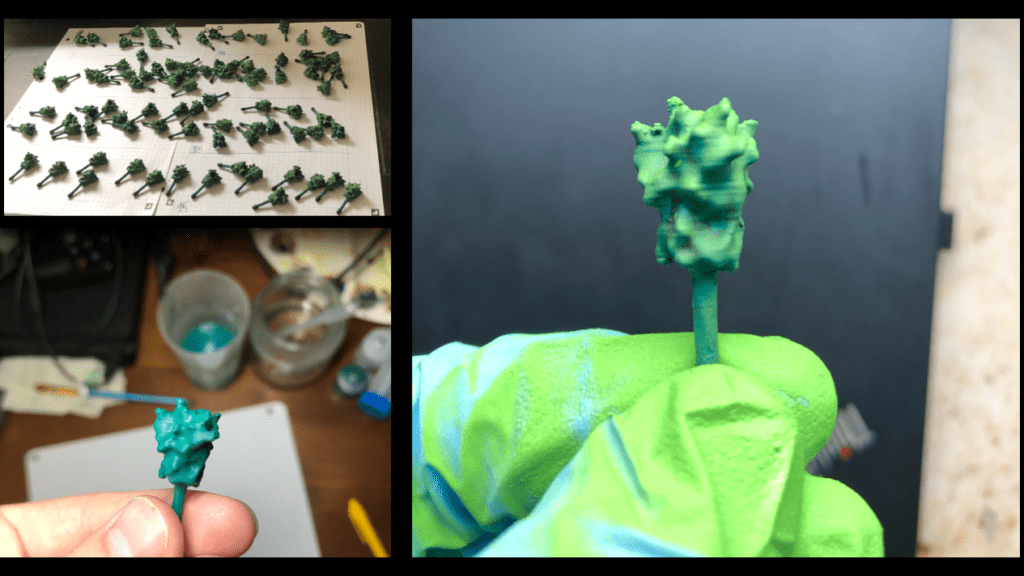

The spikes were sorted into four sets in order to produce a graded lighting affect, with those on top brighter than those lower down. If you do not plan to use a base and do not have a fixed top and bottom you can skip this part.
We highlighted each Spike Protein with a brighter lime green to achieve more contrast to create depth, which makes the surface topology easier to distinguish. Finally, the highlighting of each spike was intensified by dry-brushing the protein with the “Aqua” colour.
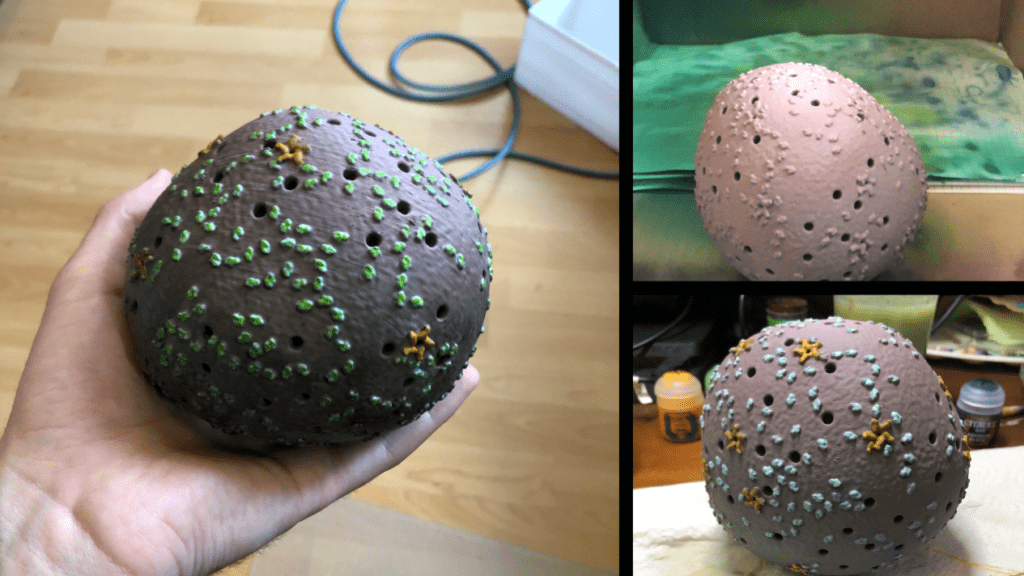

After painting was complete the spikes and virion were sealed with gloss varnish and matte finish, respectively. This step is optional; however, the varnish protects the paints against damage and wear when being handled.
Finally, the 3D model was assembled. If highlighting was used in the painting step, one should ensure the spikes are placed so that brighter spikes go on top while darker ones at the bottom. Standard modeling glue was used to hold the spikes in place, though superglue or ethyl acetate would also work fine. Because we are planning on mounting this on a stand, we have left a hole at the bottom empty where the rod of our base will go in.
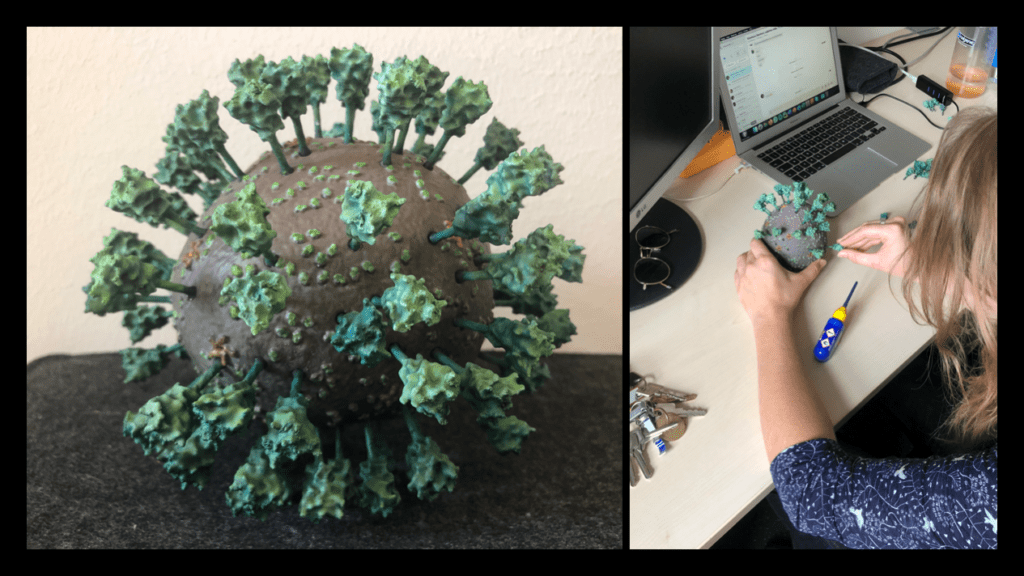

We hope that our adventure in 3D printing the Corona virus inspires you to give it a try! The process we described was completed in a little over a week. The printing jobs were completed in just over two days, the cleaning and post processing took another two days, while the painting was done over the course of a weekend. This article provides a description of our technique and should provide enough detail on how, with the outlined necessary tools, you could create a similar result. The files have been distributed through Thingiverse, and are distributed under a Creative Commons BY-NC license: You may remix, adapt, and build upon this work non-commercially and acknowledge the "Coronavirus Structural Task Force" as original author.
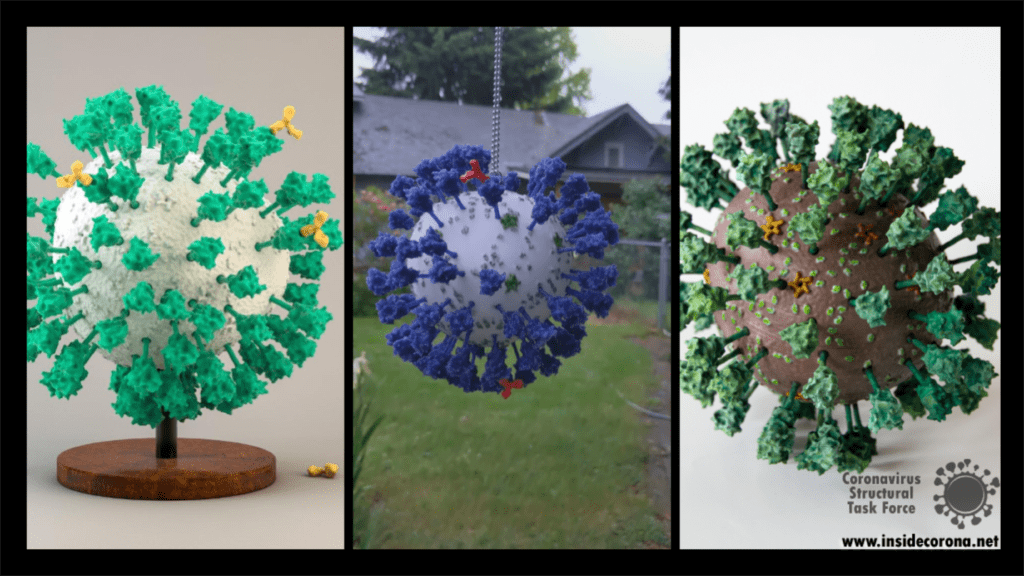

As with every 3D printed model, there are many different ways this could be tackled and achieved, and we look forward to seeing the many creative ways explored by others in this endeavor. Please do share experiences and results with us, either through the comments Thingiverse or on Twitter (you can tag us @thornlab or #insidecorona).
For a sense of perspective, we have also produced a model of the highly common rhinovirus, which is available in .stl format at the same scale as the corona virus objects. This is available at: https://www.thingiverse.com/thing:4556845.
Authors
We want to emphasize that the writing of this blog entry was a collaboration of a several people:
Dale Tronrud and Thomas Splettstoesser worked together to create the STL files for the 3D model. Dale was the person to suggest it first (with Andrea Thorn picking up on the idea). Thomas then selected the experimental models and placed all the parts to form a realistic representation. Dale provided the knowledge about the limitations imposed by the nature of 3D printing and broke up Thomas' model into printable parts that can be assembled without too much difficulty. He printed and assembled the first virion from this design.
Matt Reeves was responsible for improving the non-spherical virion model and the printing of the Würzburg model. He also determined the most suitable post-print processing techniques suitable for this project and, along with Dale and others on the team, contributed to many general technical discussions on how the model can be altered or improved further in the future.
Kristopher Nolte took part in the preprocessing and refining of the model together with Ferdinand Kirsten. Kristopher was also responsible for planning and carrying out the assembly and painting process of the Würzburg model.
An important drug target
In the first part of this series we compared the protein nsp3 from SARS-CoV and SARS-CoV-2 by sequence. Now we delve deeper into the differences between these two proteins and follow through by analyzing the structure of one domain of nsp3 in particular: papain-like protease. This domain is a very relevant drug target because of its ability not only to cleave the polyprotein, but also remove some of the post-translational modification our cells use to fight these viruses. Without papain-like protease, the virus would be unable to spread COVID-19.
Like the entire nsp3 protein, the papain-like-protease (Pl2pro) domain is localized close to the endoplasmic reticulum’s (ER) membranes. The transmembrane domains hold it in place while the majority of the protein protrudes out of the ER membrane into the cytoplasm.[1]
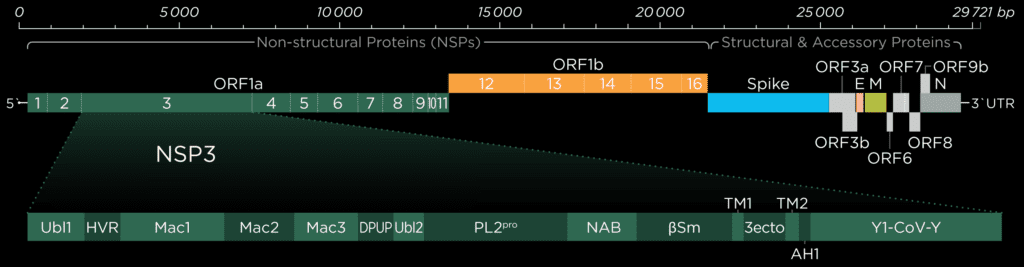
Ubiquitin-like-domain 2
We cannot discuss the Pl2pro domain without its little neighbor, which has been speculated to influence protease domain functionality.
In ubiquitin-specific proteases, the function of comparable Ubl2 domains is attributed to substrate recruitment or an increase in catalytic efficency. Ubiquitin-like-domain 2 (Ubl2) is the domain residing directly adjacent to the N-terminus of the Pl2pro catalytic domain. These ubiquitin-like domain seems to be more conserved compared to Ubl1 in different coronavirus species.[2]
If, in SARS-CoV and Murine coronavirus (MHV), Ubl2 is removed, Pl2pro loses its structural integrity. In addition, Pl2pro is then no longer able to act as an Interferon (IFN) antagonist (see below). However, some studies suggest that the Ubl2 domain in MERS-CoV might not be as essential as originally thought and in cell-based studies of this virus, Pl2pro could retain some of its enzymatic functions without the Ubl2 domain.[3]
To date, several inconsistent roles of Ubl2 were reported, and its exact function and inner workings remain enigmatic. This is being highlighted the fact that there are significant differences between the coronaviruses, and as a consequence, we need to exercise caution in applying our findings to SARS-CoV-2.
Combating the Host's Immune System
In the family of coronaviridae, viruses with either one or two Plpro domains can be found, with SARS-CoV and SARS-CoV-2 only having one. Confusingly, this single domain is however still called Pl2pro, even if it is the only papain-like protease domain in the viral genome.
Pl2pro cleaves the polyprotein from nsp1 (leader protein) up to nsp3. While Pl2pro cuts between nsp1-( ELNGG↓AV)-nsp2-( RLKGG↓AP)-nsp3-( SLKGG↓KI)-nsp4, the nsp5 (3c-like protease) cleaves the rest of the polyprotein. [2] The cysteine protease Plpro is similar to human ubiquitin-specific-protease (USP) in that it adopts a right-hand fold with "thumb", "palm" and "finger" subdomains.
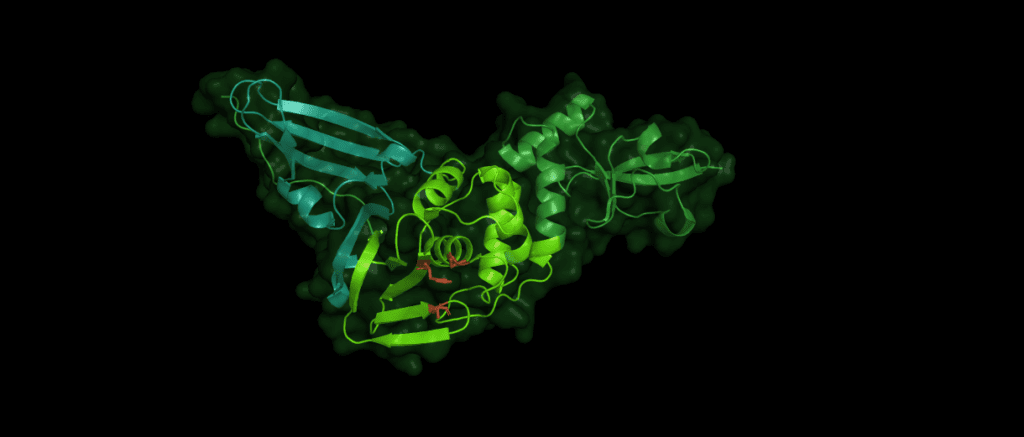
Despite the variations of Pl2pro in different coronaviridae, the same catalytic motif of three amino acid residues is essential for the stability and proteolytic activity of the domain: Cys112 is located in the thumb, His273 and Asp287 are located in the palm subdomain. (The numbers identifying these residues can vary between species.)
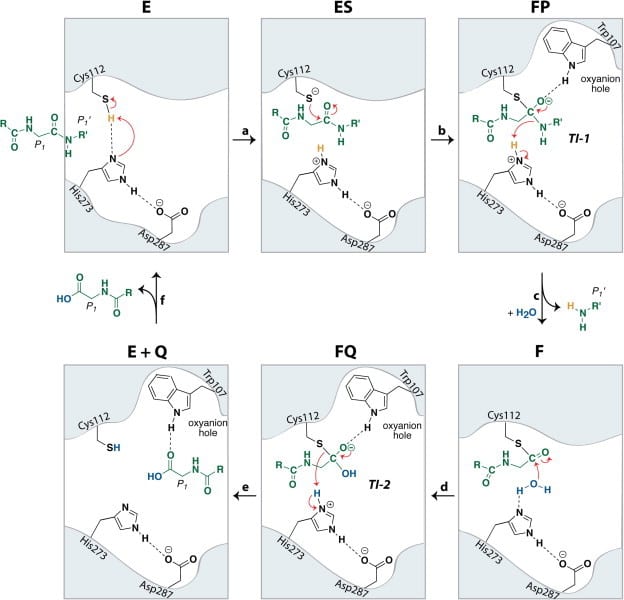
In addition, Pl2pro has deubiquitinating and deISGylating (removal of ISG15 from target proteins) abilities.[4] Both ubiquitin and ISG15 regulate facets of the immune response and through their removal Pl2pro poses as an antagonist to the human immune response. They can stimulate the production of cytokines, chemokines and other IFN-stimulated gene products which have antiviral properties. [6] ISG 15 is an ubiquitin-like modifier composed of two ubiquitin-like folds that has an essential role in marking newly synthesized proteins during the antiviral response.[3] Post-translational modification by ubiquitin and interferon-stimulating gene 15 (ISG15) is reversed by isopeptide bond hydrolysis. Figure 3 shows a proposed mechanism for the cleaving of isopeptide bonds by SARS-CoV.
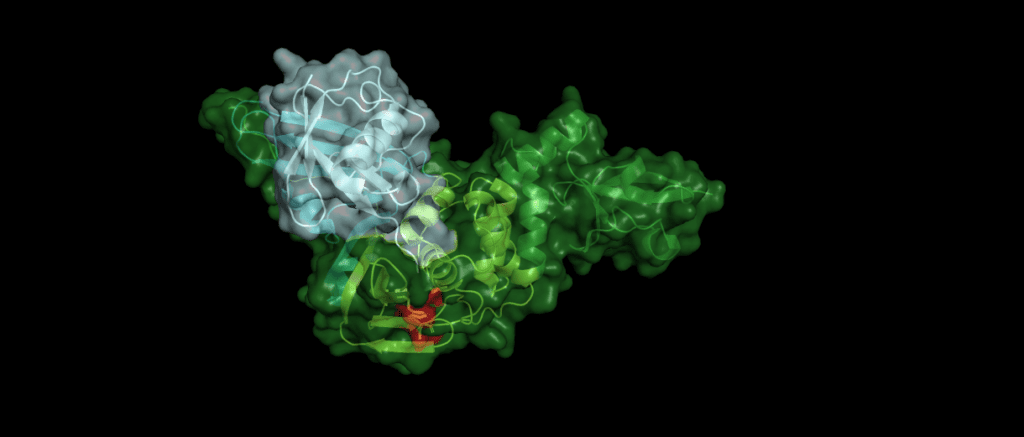
An example
Toll-like receptors (TLRs) are an important part of the machinery of the human immune response, which recognizes the pathogen-associated molecular patterns. The ability of the host cell to transduce the so-called Toll-like receptor 7 (TLR7) mediated immune response is diminished (Fig. 5) by Pl2pro as it removes Lys63-linked-ubiquitin from the TNF receptor associated factors TRAF3 and TRAF6. [5]
In addition, SARS-CoV can hamper the antiviral activities of interferon. The Pl2pro domain inhibits in combination with a transmembrane (TM) domain the STING mediated activation of interferon expression. PL2pro-TM interacts with TRAF3, TBK1, IKKε, STING and IRF3, the key components assembling a regulatory complex for activation of IFN expression.[5]
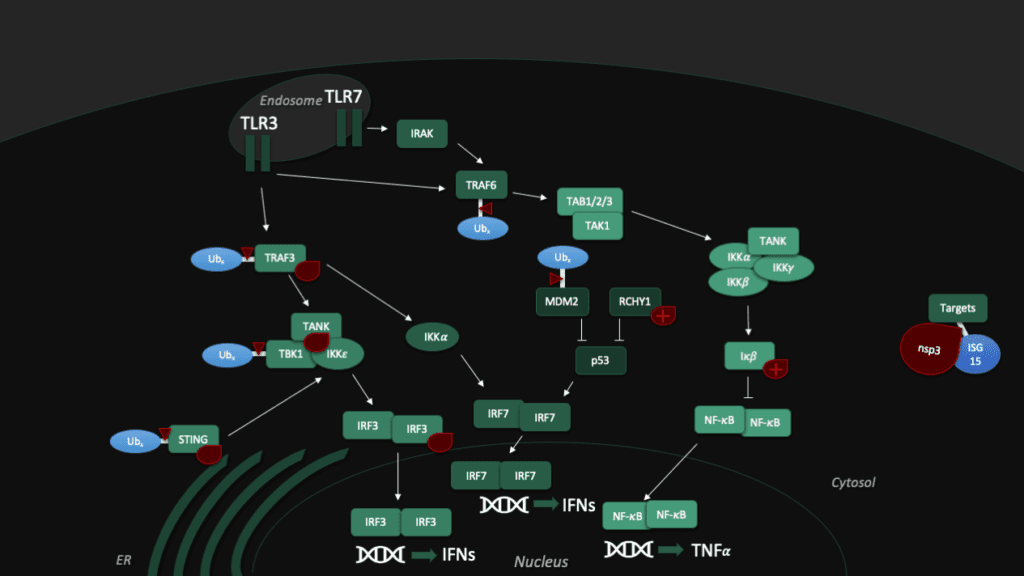
Another tool to fight the coronavirus in human cells is the "guardian of the genome", p53. The tumor supressor protein p53 impedes the replication of SARS-CoV, though the virus fights back with Pl2pro, which binds a p53 degradation stimulator named "RING finger and CHY zinc finger domain-containing protein 1" (or short: RCHY1). Enhanced by the Macro somains in NSP3, this binding enhances the stability of RCHY1 and hence promotes the degradation of p53. In addition, Pl2pro blocks another crucial cellular defense mechanism: The NF-κB pathway, which regulates immune responses to infections. SARS-CoV Pl2pro can stabilize IκBα, an inhibitor of NF-κB.[3]
Although all Pl2pro in different coronaviridae suppress the immune response, the targets differ between various species. For example, SARS-CoV Pl2pro preferentially processes Lys48 linked poly-ubiquitin chains, which are markers for proteasome degradation. MERS, on the other hand, shows no differences in effectivity between Lys48 and Lys63 linked di-Ubq chains. Lys63-linked chains are related to signal transduction cascades of the host immune system. Studies have shown that specificity among Pl2pro for Ubiquitin and ISG15 substrates can be altered with as little as a single amino acid change.[6] However, even though there are differences, for SARS-CoV-2, it is likely that at least some of the functions are similar.
Structural comparison
In order to predict Pl2Pro function for the novel Coronavirus SARS-CoV-2, we start by aligning their sequence like we did in the first part of this series to comapare the sequence with the one from SARS-CoV-2. Both domains share a similarity of 82.8% over the length of 313 amino acids. However, this time, we go for a more detailed analysis of the 54 individual differences, which are:
T3R N14I V20V N48N H49S V56Y D60N E66V D75T S77P P95Y G99N S114A V115T L116A L119T E123I K125L P129P A134D A143E N155C H170S L171Y Q173F S179D K181C C191T T195Q T196Q G200K N214E L215Q K216F G218K I221Q C225T D228K A229Q Y232K F240P Y250Q L252E Q254K G255H C259T E262S H274K K278S I284C L289L S293S T300I S308N
(The first letter refers to SARS-CoV, and the second to the amino acid residue in SARS-CoV-2.)
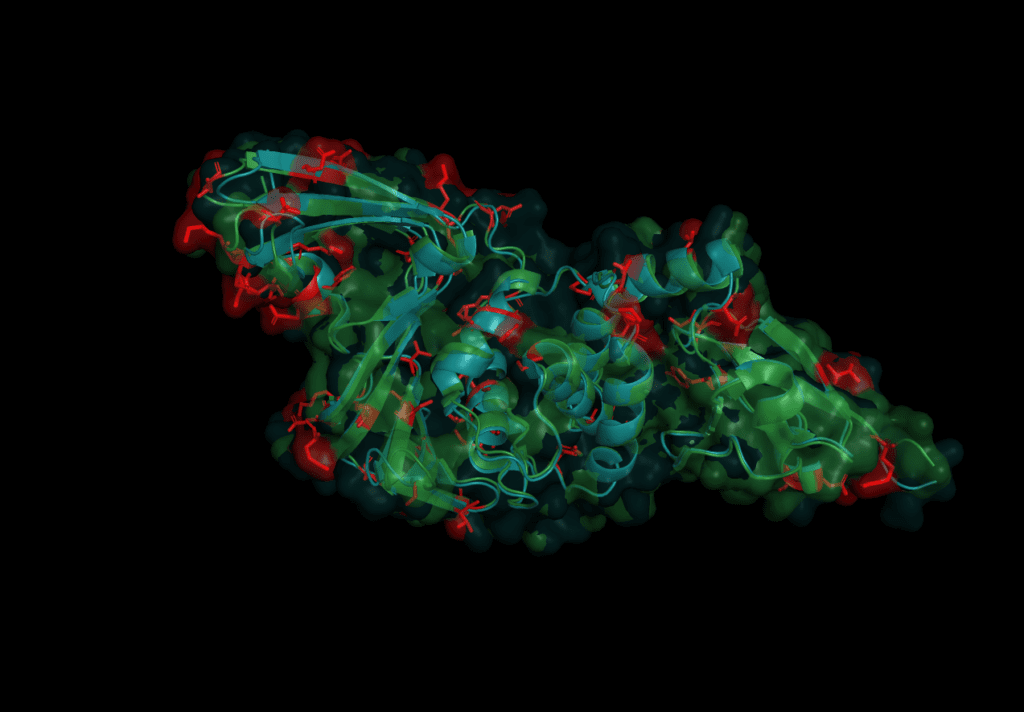
The mutations are evenly spread over the protein. None of the catalytic triad (Cys 112, His 273, Asp287) are changed as is to be expected given their conservation in all other coronaviruses. On further investigation, however, in the motif which interacts with ubiquitin six sites are different: S170T, Y171H, F216L, Q195K, T225V, and K232Q. Earlier studies concluded that the mutation of position 232 from Glutamine to Lysine increases the affinity for ubiquitin at the expense of the de-ubiquitination effectiveness.[6] The kinetics of SARS-CoV-2 nsp3 Pl2pro were studied to test if the protease domain of nsp3 has a reduced effectiveness in binding ubiquitin compared to nsp3 from SARS-CoV, MERS-CoV.
All three Pl2Pro variants cleave more ISG15 than ubiquitin. SARS-CoV has the fastest kinetics of the three viruses. And, the slower kinetics of SARS-CoV-2 resemble those of MERS-CoV rather more than SARS-CoV, having a 10 times higher turnover rate (kcat) as a deISGylase than as a deubiquitase.[6]
Besides the kinetics, the Pl2pro’s affinity for different poly-ubiquitin linkage sites was measured. The result shows that while SARS-CoV-2 can cut K48-Ub linked polyproteins, it seems to lack an ability to cut other polyubiquitin chains. Those K48-Ub linked polyproteins are cleaved at a slower rate than by SARS-CoV. In this regard, SARS-CoV-2 distinguishes itself from MERS-CoV which has the ability to cleave K63-linkages. It is suggested that the decrease in deubiquitinase effectiveness may not be irrelevant, but could lead to the often-mild symptoms that are a factor in why SARS-CoV-2 has been able to evade our efforts in quarantine. But this is mere speculation and a lot more research is needed to resolve the matter.[6]
PL2pro as a drug target
Pl2pro was a potential drug target early on in SARS-CoV-2 research. Hilgenfeld et al. name two major challenges we have to overcome to find a drug targeting Pl2pro. One is that the binding sites are tailor-made to bind glycine residues. Also, this very specific binding motif is rather ubiquitious in our cells. These two problems make it difficult to find an inhibitor which fits and is specific to Pl2pro. However, scientists found a weak spot: a loop called Blocking Loop 2 (BL2) regulates substrate binding and may be a promising target to inhibit PL2pro.[2] Naphthalene based inhibitors, which were earlier proposed to inhibit the BL2 of SARS-CoV, were shown to also inhibit SARS-CoV-2 Pl2pro, in particular an inhibitor called GRL-0617.[6]
For in-silico drug development, it might be prudent to choose high-resolution structures which already have a ligand or inhibitor bound, such as 6yva, 6wuu, 6wx4 or 6yaa. Technically speaking, 6wrh, albeit being a mutant, is one of the highest-quality structures available for SARS-CoV-2 Pl2pro.
In fact, a lot of research is still required to consolidate our understanding of this protein and its domains. In spite of that, we are making progress in our endeavor to fight this virus - and every step we take is one more to win this fight.
Sources
[1] Báez-Santos YM, St John SE, Mesecar AD. The SARS-coronavirus papain-like protease: structure, function and inhibition by designed antiviral compounds. Antiviral Res. 2015;115:21-38. doi:10.1016/j.antiviral.2014.12.015, https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5896749/
[2] Lei J, Kusov Y, Hilgenfeld R. Nsp3 of coronaviruses: Structures and functions of a large multi-domain protein. Antiviral Res. 2018;149:58-74. doi:10.1016/j.antiviral.2017.11.001, https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7113668/
[3] Clasman JR, Báez-Santos YM, Mettelman RC, O'Brien A, Baker SC, Mesecar AD. X-ray Structure and Enzymatic Activity Profile of a Core Papain-like Protease of MERS Coronavirus with utility for structure-based drug design. Sci Rep. 2017;7:40292. Published 2017 Jan 12. doi:10.1038/srep40292, https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5228125/
[4] Lei J, Hilgenfeld R. RNA-virus proteases counteracting host innate immunity. FEBS Lett. 2017;591(20):3190-3210. doi:10.1002/1873-3468.12827, https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7163997/
[5] Chen X, Yang X, Zheng Y, Yang Y, Xing Y, Chen Z. SARS coronavirus papain-like protease inhibits the type I interferon signaling pathway through interaction with the STING-TRAF3-TBK1 complex. Protein Cell. 2014;5(5):369-381. doi:10.1007/s13238-014-0026-3, https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3996160/
[6] Freitas BT, Durie IA, Murray J, et al. Characterization and Noncovalent Inhibition of the Deubiquitinase and deISGylase Activity of SARS-CoV-2 Papain-Like Protease [published online ahead of print, 2020 Jun 4]. ACS Infect Dis. 2020;acsinfecdis.0c00168. doi:10.1021/acsinfecdis.0c00168, https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7274171/
