A guest entry by Hauke Hillen
In order for the novel coronavirus SARS-CoV-2 to replicate, it has to achieve two basic tasks: It needs to make copies of its genome that can be packaged into new virus particles, and it needs to activate viral genes to produce the proteins that actually form new virus particles, such as spike or nucleocapsid. Both tasks are carried out by a specialized molecular copying machine called the replication and transcription complex, or short RTC. The RTC is made up of a number of viral non-structural proteins (nsps) which act together to produce copies of the viral RNA genome. Some of the RTC components have been discussed in previous posts, for example the exonuclease nsp14, which can correct errors that occur during RNA copying (this is called proofreading),or the methyltransferases nsp14 and nsp16 that add chemical modifications to the RNA that help stabilize and hide it from the immune system (this is called capping).
In this post, we will have a closer look at the enzyme that carries out RNA replication, the RNA-dependent RNA polymerase (RdRp) nsp12, and how it interacts with the other components to form the RTC.
RNA polymera…what?
First off, let’s briefly discuss what a “RNA-dependent RNA polymerase” is, and why it is important for the virus. Polymerases are enzymes found in every living cell carrying out one of the most fundamental tasks in biology: they replicate genetic information. While most cells store their genetic information in form of DNA (deoxyribonucleic acid) and use RNA (ribonucleic acid) only as transient messenger molecules (mRNAs), some viruses rely on RNA for both information storage and transmission and are hence called “RNA viruses”. To replicate their genetic information, they need a polymerase that uses RNA as template to copy the encoded information into a new RNA molecule – a RNA-dependent RNA polymerase. Chemically speaking, RNA is a polymer (a long string of nearly-identical individual building blocks) composed of the four nucleotides adenosine (A), guanosine (G), cytosine (C) and uracil (U), the sequence of which defines the genetic information.
The job of the RNA polymerase is to read the sequence of nucleotides in a template RNA and synthesize new RNA with the same sequence (technically with a complementary sequence) using individual nucleotides as building blocks. To do this, the RNA polymerase has to achieve three basic steps. First, it needs to bind the template RNA. Second, it has to read the sequence of nucleotides in the RNA. Third, it has to incorporate the correct matching nucleotide building blocks to polymerize a new RNA strand.
Coronaviruses are RNA viruses and therefore have an RNA-dependent RNA polymerase (RdRp, or nsp12). However, they are exceptional in several ways. First, their RNA genomes are almost 30.000 nucleotides in length, which are the largest RNA virus genomes known to date. Second, their RNA polymerase nsp12 requires the additional proteins nsp7 and nsp8 in order to form the active “core” RdRp. Third, this core RdRp assembles with further viral proteins to form the RTC, which can carry out additional functions such as proofreading to remove copying errors - a highly unusual capability for RNA viruses.
Since the RNA polymerase has such a fundamental job during virus replication, it is an attractive drug target to combat viral infections. Indeed, many successful anti-viral drugs against Hepatitis C virus, HIV or Herpes virus act by inhibiting viral polymerases. Strikingly, viral RdRp enzymes are remarkably similar in their overall structure even between unrelated viruses, indicating that they share a common evolutionary ancestor and their function is so essential that it does not allow for drastic changes. Therefore, some known anti-virals developed to treat other viral diseases have also been tested for their activity against the polymerase of SARS-CoV-2 and even approved for clinical use against COVID-19 [1]. However, these repurposed compounds are generally not as effective as many had hoped, because even rather subtle structural differences between the polymerase enzymes of different viruses can have strong effects on the action of anti-viral drugs. Thus, more specific and efficient drugs against SARS-CoV-2 are therefore badly needed. In order to discover and develop such compounds, detailed knowledge of the structure and function of the RdRp is necessary.
Structure of the coronavirus RNA-dependent RNA polymerase (RdRp)
The first structure of a coronavirus polymerase was determined shortly before the outbreak of the current COVID-19 pandemic when Kirchdoerfer and Ward reported the cryo-electron microscopy (cryo-EM) structure of SARS-CoV-1 RdRp [2]. Since SARS-CoV-2 has emerged, scientists all over the world have been racing to determine the structures of its RdRp. As of February 2021, this has led to more than 20 structures of SARS-CoV-2 polymerase-complexes published in the PDB, and often several groups of scientists reported similar structures around the same time.
These structures show that the RNA polymerase nsp12 resembles a right hand with individual domains called palm, fingers and thumb (Figure 1) [3–7]. This “hand” shape is typical for viral RNA polymerases and holds a tight grip on the double-stranded RNA helix that forms between template and product strand during RNA synthesis. Within the palm lies the “active center” of the enzyme, where nucleotides are added to the growing product chain. The active center is accessible from the surface of the enzyme through a special tunnel, so that nucleotides can enter the substrate-binding site. As the template strand is opposite to the substrate-binding site, each nucleotide entering is “sampled” for whether it can form base-pairing interactions with the template base. If this is the case, the nucleotide remains bound, and it is added to the 3’ end of the product strand by forming a chemical bond. After that, the RdRp enzyme must slide ahead on the template strand by one nucleotide, which moves the newly produced 3’ end of the product RNA from the substrate-binding site (which is sometimes also referred as position +1) to the position where the previous 3’ end of the product was located prior to addition (position -1). This “translocation” completes the nucleotide addition cycle, as it positions the next templating base and frees up the substrate-binding site for the next matching building block.
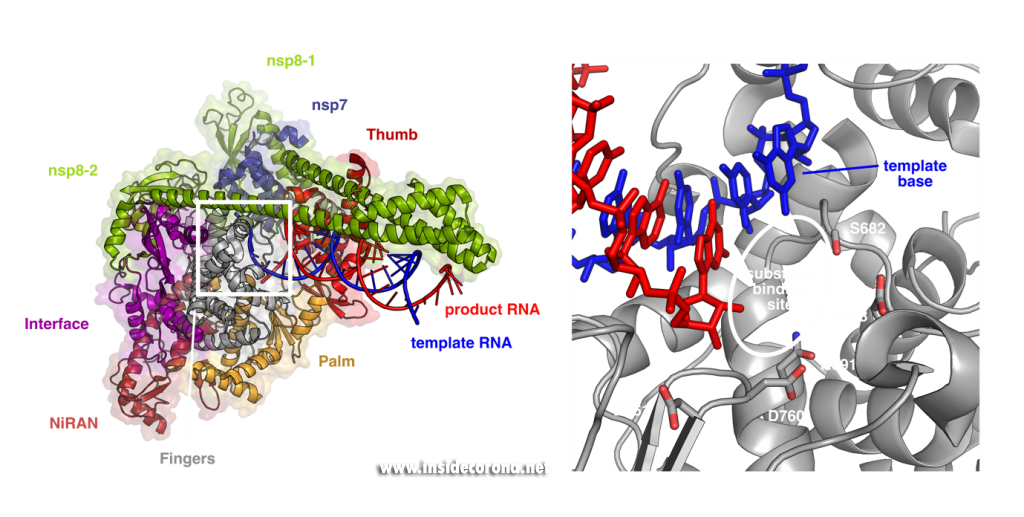
Left: Cryo-EM structure of SARS-CoV-2 RdRp (PDB 6YYT). Right: Enlarged active site with template strand in blue and nascent chain in red..
In addition to its polymerase domain, RdRp also contains a part that is only found in nidoviruses (the virus family that coronaviruses belong to) called “nidovirus RdRp-associated nucleotidyltransferase domain”, or short NiRAN-domain. Scientists believe that this domain has the capability to transfer nucleotidyl-residues, which means that it can form chemical bonds between nucleotides and other molecules. This hints that it may be involved in modification of the RNA (so-called “capping”) or in helping the enzyme initially kickstart RNA synthesis, but its precise role during coronavirus replication is still being studied by scientists.
In order to efficiently copy RNA, nsp12 requires two additional viral proteins, nsp7 and nsp8. The structures of coronavirus RdRp show that two molecules of nsp8 and one molecule of nsp7 bind on top of the hand-shaped nsp12. Interestingly, even though identical in amino acid sequence, the two nsp8 molecules adopt slightly different shapes. While one of them interacts with the finger domain of nsp12 directly, the interaction of the other one is mediated by nsp7. Both nsp8 molecules have long “arms” that protrude away from the polymerase and touch the RNA duplex as it emerges from the polymerase during replication. These “sliding poles” are unique to coronaviruses and most likely stabilize the RdRp on the RNA, which may help to make sure it doesn’t fall off during replication of the very large genome.
Seeing is believing - visualizing how anti-viral compounds block RdRp
So how exactly can this structural knowledge help to find new drugs against COVID-19? In many ways, enzymes are like tiny molecular machines. By studying their structure, one can analyze in detail how they work biochemically, and this in turn allows us to come up with ways to block their function. Most known anti-viral drugs that target RNA polymerases are so-called nucleoside analogs, which means they are molecules that structurally resemble the natural building blocks of RNA. These compounds can “trick” the RdRp by binding to the active site, but due to their chemical nature, they either cannot be incorporated into the product or lead to mutations that end the viral life cycle. The structures of SARS-CoV-2 RdRp reveal the exact architecture of the active site and show how the chemical environment that the enzyme creates around the product RNA and the substrate nucleotides, facilitates polymerization (Figure 1). This knowledge can help to rationally design or improve compounds in such a way that they bind more efficiently.
This is exemplified by recent studies analyzing how repurposed anti-virals inhibit SARS-CoV-2 RdRp. One such drug is Remdesivir, a compound originally developed against Ebola and other viruses that has also been approved by the FDA and European agencies for treatment of COVID-19 (see also this previous post). Remdesivir chemically resembles adenosine triphosphate (ATP), but has an additional bulky chemical residue called a cyano-group attached to the C1-atom of its ribose moiety. In contrast to most nucleoside analogs, Remdesivir does not block the RdRp immediately, but only after another three nucleotides are added, a process called “delayed stalling” [8–10]. Initial structures of SARS-CoV-2 RdRp in the presence of Remdesivir showed how it can act as an adenosine analog and how it can be incorporated at the 3’ end of the nascent product RNA strand and translocated to the -1 position (Figure 2a,b) [5,6]. However, this could not explain how it would lead to inhibition of RNA synthesis.
To pinpoint why Remdesivir interferes with RNA synthesis exactly after three subsequent nucleotides are added, the authors of a recent study used a combination of synthetic chemistry and structural biology [11]. They systematically determined structures of SARS-CoV-2 RdRp bound to a template-product RNA duplex which contained Remdesivir and either two or three additional nucleotides at the 3’ end. In the first case, the structure showed that the RdRp was in the post-translocated state, with Remdesivir at position -3 and an empty substrate binding site, as expected (Figure 2c). In contrast, the structure of the RdRp with an RNA containing Remdesivir and three additional nucleotides was not in the post-translocated state and Remdesivir was not located at position -4. Instead, it remained at position -3, and the third additional nucleotide at the 3’ end of the product was stuck in the substrate-binding site (Figure 2d). This state resembles the situation directly after addition of a new nucleotide but before translocation and is hence called the pre-translocated state. This suggests that remdesivir inhibits the SARS-CoV-2 RNA polymerase by posing a translocation barrier, and the structures provide a molecular and chemical explanation for this: Initially, Remdesivir can be added to the growing RNA just like adenosine triphosphate and also translocated to add another two nucleotides. However, after binding and addition of a third nucleotide, Remdesivir can not be translocated to the -4 position, because its bulky cyano group would clash with a serine residue (Ser861) in the thumb domain of nsp12 (Figure 2c). Therefore, the polymerase gets stuck in the pre-translocated state, which explains why exactly three nucleotides can be added after Remdesivir incorporation – addition of a fourth nucleotide would first require translocation. This proposed mechanism is in agreement with previous modelling [5,10] and was shortly after independently confirmed by another structural study, in which the authors managed to trap an identical pre-translocated, stalled intermediate with Remdesivir in position -3 [12].
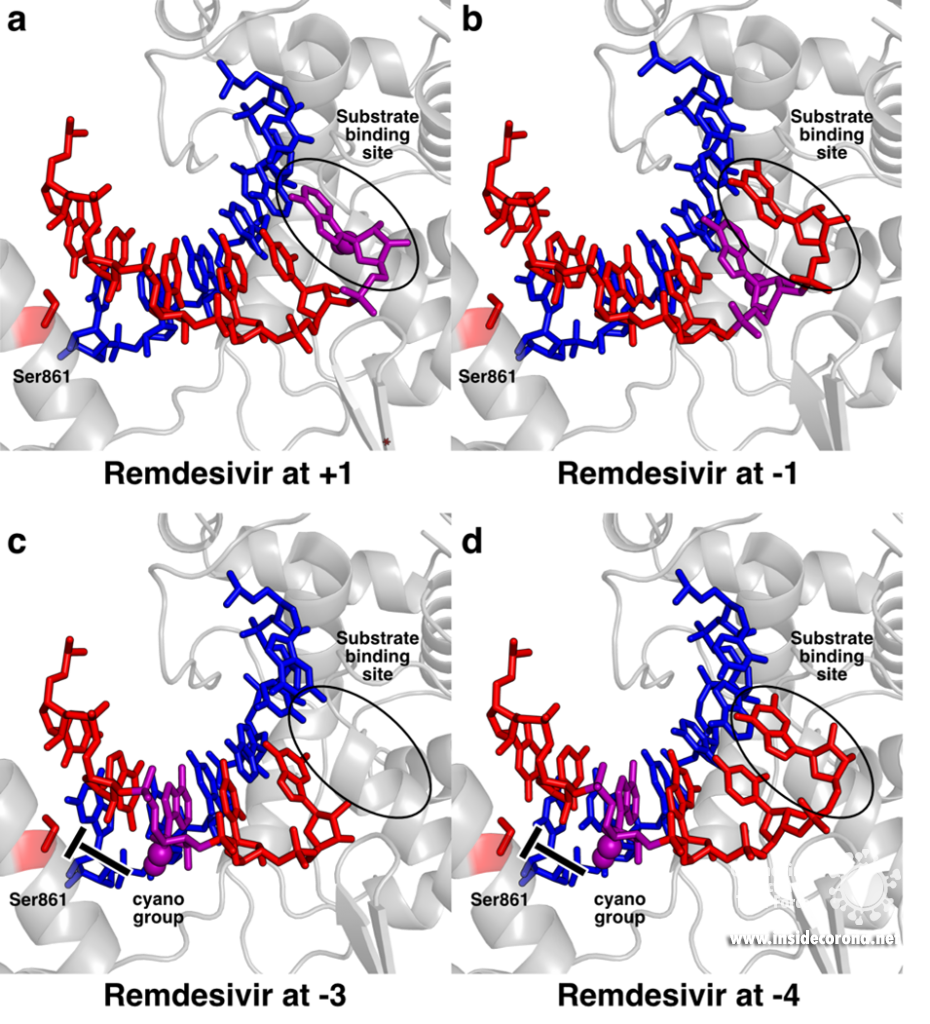
Structural snapshots of remdesivir (purple) moving through the active site of SARS-CoV-2 RdRp. When it reaches the third position after its incorporation to the RNA (-3), its further movement is blocked because the cyano group would bump into Ser861. A) Remdesivir at position +1 (PDB 7BV2) B) Remdesivir at position -1 PDB 7C2K C) Remdesivir at position -3 (PDB 7B3B) D) Remdesivir at position -3 with the nucleotide at the 3’ end stuck in the substrate binding site (PDB 7B3C).
This mechanism also suggests how Remdesivir may at least partially escape the coronavirus proofreading enzyme nsp14, which removes misincorporated nucleotides at the 3’ end of the RNA and thus counteracts anti-virals that target RdRp. Since Remdesivir can be translocated until it reaches position -3 before it causes stalling, it may leave the active site of the enzyme before it can be recognized by the proofreading machinery.
Importantly, these studies also provide clues as to why Remdesivir has had limited success in fighting COVID-19. The structures show that the steric block between Ser861 and the cyano group of Remdesivir is not severe and can therefore be overcome by the enzyme, for example at high concentrations of substrate NTPs [8] Consistent with this, substitution of Ser861 with residues that clash even less (Alanine or Glycine) make the RdRp less sensitive or even resistant to Remdesivir [5,13]. This suggests that the translocation barrier could potentially be enhanced by a compound that leads to more severe clashes. One way to achieve this could be to modify Remdesivir to contain more bulky chemical moieties than the cyano group. Thus, the detailed molecular insights into the mechanism of Remdesivir also provide a rational basis for designing more potent anti-virals and test their effect on the SARS-CoV-2 RdRp.
Similar structure-function studies are now being undertaken also for other promising anti-viral compounds, such as Favipiravir. Like Remdesivir, it is a nucleoside analogue that was initially developed against other viruses, but showed some promising results against SARS-CoV-2. Structures of the SARS-CoV-2 RNA polymerase with Favipiravir show how it mimics both guanosine or adenosine in the active site of the enzyme by forming unusual base-pairing interactions with cytosine and uracil, respectively, and this leads to errors during RNA copying that eventually kill the virus [14,15]. Another study recently reported the structure of Suramin bound to SARS-CoV-2 RdRp [16]. In contrast to Remdesivir and Favipiravir, this compound is not a nucleoside analog and hence does not get incorporated into the RNA. Instead, two Suramin molecules can apparently bind to the RdRp and thereby prevent its association with template and product RNA, rendering it inactive.
These studies are good examples of how structural biology can visualize complicated chemical reactions in an intuitive way. Based on these results, drugs like Remdesivir or Favipiravir can be rationally optimized to more effectively combat COVID-19 and may cause less side effects.
Dissecting the SARS-CoV-2 RTC structure by structure
In addition to provide detailed snapshots of how anti-viral compounds act to inhibit SARS-CoV-2 RdRp, structural studies are also helping scientists to understand how the unique coronavirus RTC combines different functions such as RNA synthesis, proofreading and modification. After the structure of the “core” SARS-CoV-2 RdRp was determined within a few months after the outbreak of COVID-19, scientists quickly moved to studying how additional non-structural proteins bind to it to form the RTC. One of these is nsp13, which belongs to a protein class called “helicases”. These are enzymes that bind to DNA or RNA and, with the help of chemical energy in the form of ATP, move along them or unwind helices. Cryo-EM structures of nsp13 bound to the SARS-CoV-2 RdRp show that two molecules of nsp13 can bind to the RdRp, and suggest that it may allow the polymerase to move backwards on the RNA (Figure 3) [17,18]. This finding seems unexpected at first, but experts think this may be required for the proofreading enzyme nsp14 to remove errors that the polymerase makes during copying or to produce the mRNAs for certain viral proteins. Another recent cryo-EM structure shows that the small protein nsp9 interacts with the NiRAN domain in the SARS-CoV-2 polymerase [19]. In the accompanying paper, the authors suggest that the NiRAN domain may be involved in RNA capping, and nsp9 seems to block its activity. However, others have proposed that nsp9 is in fact a substrate of nucleotidylation by the NiRAN domain, and that it may be involved in priming the RNA polymerase for initial RNA synthesis [20]. Thus, further studies are necessary to determine whether the NiRAN is involved in capping, priming or even both.
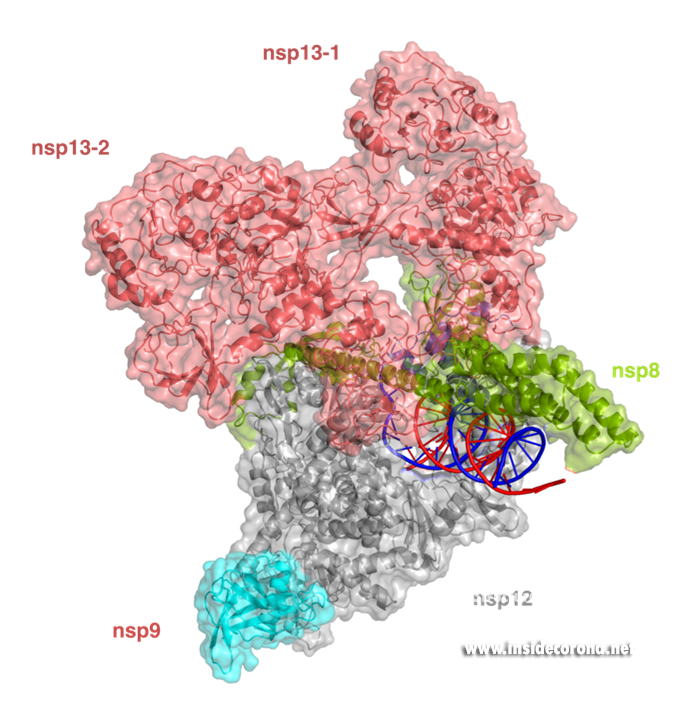
Structure of the SARS-CoV-2 RdRp complex with nsp13 (salmon) and nsp9 (cyan) (PDB 7CYQ).
What’s next?
Over the past year, scientists have uncovered the structure of the SARS-CoV-2 RdRp and associated proteins at a record-breaking pace. While these structures provide impressive first glimpses at RdRp-complexes, researchers are already working to determine how the remaining nsp proteins interact with the RdRp to form the complete RTC complex. This may ultimately aid the quest to find new treatment options for COVID-19 that not only target the polymerase itself, but also proofreading or RNA capping. Such drugs are not only desperately needed for the current pandemic, but may also prove useful for future emerging coronaviruses, because the RNA polymerase is typically very similar even between different virus strains.
1. Ledford H: Hopes rise for coronavirus drug remdesivir. Nature 2020, doi:10.1038/d41586-020-01295-8.
2. Kirchdoerfer RN, Ward AB: Structure of the SARS-CoV nsp12 polymerase bound to nsp7 and nsp8 co-factors. Nat Commun 2019, 10:2342.
3. Gao Y, Yan L, Huang Y, Liu F, Zhao Y, Cao L, Wang T, Sun Q, Ming Z, Zhang L, et al.: Structure of the RNA-dependent RNA polymerase from COVID-19 virus. Science 2020, 368:779–782.
4. Hillen HS, Kokic G, Farnung L, Dienemann C, Tegunov D, Cramer P: Structure of replicating SARS-CoV-2 polymerase. Nature 2020, 584:154–156.
5. Wang Q, Wu J, Wang H, Gao Y, Liu Q, Mu A, Ji W, Yan L, Zhu Y, Zhu C, et al.: Structural Basis for RNA Replication by the SARS-CoV-2 Polymerase. Cell 2020, 182:417-428.e13.
6. Yin W, Mao C, Luan X, Shen D-D, Shen Q, Su H, Wang X, Zhou F, Zhao W, Gao M, et al.: Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir. Science 2020, 368:1499–1504.
7. Peng Q, Peng R, Yuan B, Zhao J, Wang M, Wang X, Wang Q, Sun Y, Fan Z, Qi J, et al.: Structural and Biochemical Characterization of the nsp12-nsp7-nsp8 Core Polymerase Complex from SARS-CoV-2. Cell Reports 2020, 31:107774.
8. Gordon CJ, Tchesnokov EP, Woolner E, Perry JK, Feng JY, Porter DP, Götte M: Remdesivir is a direct-acting antiviral that inhibits RNA-dependent RNA polymerase from severe acute respiratory syndrome coronavirus 2 with high potency. J Biol Chem 2020, 295:6785–6797.
9. Tchesnokov EP, Feng JY, Porter DP, Götte M: Mechanism of Inhibition of Ebola Virus RNA-Dependent RNA Polymerase by Remdesivir. Viruses 2019, 11:326.
10. Gordon CJ, Tchesnokov EP, Feng JY, Porter DP, Gotte M: The antiviral compound remdesivir potently inhibits RNA-dependent RNA polymerase from Middle East respiratory syndrome coronavirus. The Journal of biological chemistry 2020, doi:10.1074/jbc.ac120.013056.
11. Kokic G, Hillen HS, Tegunov D, Dienemann C, Seitz F, Schmitzova J, Farnung L, Siewert A, Höbartner C, Cramer P: Mechanism of SARS-CoV-2 polymerase stalling by remdesivir. Nat Commun 2021, 12:279.
12. Bravo JPK, Dangerfield TL, Taylor DW, Johnson KA: Remdesivir is a delayed translocation inhibitor of SARS CoV-2 replication in vitro. Biorxiv 2020, doi:10.1101/2020.12.14.422718.
13. Tchesnokov EP, Gordon CJ, Woolner E, Kocinkova D, Perry JK, Feng JY, Porter DP, Götte M: Template-dependent inhibition of coronavirus RNA-dependent RNA polymerase by remdesivir reveals a second mechanism of action. J Biol Chem 2020, 295:16156–16165.
14. Naydenova K, Muir KW, Wu L-F, Zhang Z, Coscia F, Peet MJ, Castro-Hartmann P, Qian P, Sader K, Dent K, et al.: Structural basis for the inhibition of the SARS-CoV-2 RNA-dependent RNA polymerase by favipiravir-RTP. Biorxiv 2020, doi:10.1101/2020.10.21.347690.
15. Peng Q, Peng R, Yuan B, Wang M, Zhao J, Fu L, Qi J, Shi Y: Structural basis of SARS-CoV-2 polymerase inhibition by Favipiravir. Innovation 2021, doi:10.1016/j.xinn.2021.100080.
16. Yin W, Luan X, Li Z, Zhou Z, Wang Q, Gao M, Wang X, Zhou F, Shi J, You E, et al.: Structural basis for inhibition of the SARS-CoV-2 RNA polymerase by suramin. Nat Struct Mol Biol 2021, 28:319–325.
17. Chen J, Malone B, Llewellyn E, Grasso M, Shelton PMM, Olinares PDB, Maruthi K, Eng ET, Vatandaslar H, Chait BT, et al.: Structural basis for helicase-polymerase coupling in the SARS-CoV-2 replication-transcription complex. Cell 2020, doi:10.1016/j.cell.2020.07.033.
18. Yan L, Zhang Y, Ge J, Zheng L, Gao Y, Wang T, Jia Z, Wang H, Huang Y, Li M, et al.: Architecture of a SARS-CoV-2 mini replication and transcription complex. Nat Commun 2020, 11:5874.
19. Yan L, Ge J, Zheng L, Zhang Y, Gao Y, Wang T, Huang Y, Yang Y, Gao S, Li M, et al.: Cryo-EM Structure of an Extended SARS-CoV-2 Replication and Transcription Complex Reveals an Intermediate State in Cap Synthesis. Cell 2021, 184:184-193.e10.
20. Slanina H, Madhugiri R, Bylapudi G, Schultheiß K, Karl N, Gulyaeva A, Gorbalenya AE, Linne U, Ziebuhr J: Coronavirus replication–transcription complex: Vital and selective NMPylation of a conserved site in nsp9 by the NiRAN-RdRp subunit. Proc National Acad Sci 2021, 118:e2022310118.
The building plan
Storing the building plans for a virus in its genome is much like how we store ideas in language. This may sound strange but, as an example, typos in spelling, grammar, or word usage, can lead to the meaning of a sentence either changing dramatically, remaining virtually unchanged, or becoming complete nonsense. The SARS-CoV-2 genome consists of RNA. Transcription of this RNA runs into a similar problem: errors can lead to the loss of function, a gain of function, or be completely inconsequential to the resulting protein (Figure 1). Large changes may break the virus, but smaller changes may provide an advantage and are essential for evolution.
Targeting the copy machine
In a previous article we spoke about the copy machinery of the virus, including the RNA-dependent RNA polymerase (RdRp), and drugs targeting it, such as Remdesivir. The goal of these drugs is to jam the enzyme and halt RNA production - or to cause more errors than are sustainable, with the end result being a less infectious virus. The reason the development of drugs targeting the copy machinery of RNA is worthwhile is that humans don’t have machinery to reproduce RNA from RNA. This means drugs targeting this machinery are less likely to interfere with normal processes in people. What if the virus could quickly repair these errors before the new genome is packed into a hull and kicked out the door? That would make finding a therapeutic much more difficult…
Correctional facilities
Unfortunately, SARS-CoV-2 has a way to repair the mistakes. When errors are introduced in transcription through environmental mutagenesis or even mutations caused by nucleotide analogs like Ribavarin1–3, the non-structural protein 14 (nsp14) has the ability to remove them. This multifunctional protein removes errors with the exoribonuclease (ExoN) activity of its N-terminal domain, while the C-terminal domain has the unrelated function of methylating the end cap of the viral RNA3,4.
However, this ExoN does not work alone. There is a replication complex made up of proteins performing many roles in the production of new RNA with high fidelity. Nsp12 is the main hub that makes a new RNA chain to complement the template. Nsp7 and nsp8 have a “processivity” role to enable nsp12 to function efficiently. In addition to these proteins there is a two-component proofreading system of Helicase (nsp13) and the ExoN domain of nsp14. Helicase can detect misshapen RNA helices caused by errors made by the copy machinery5. It then unwinds these double strands of RNA and feeds the strand containing the error into the ExoN domain of nsp14 where they are chopped out. This results in nsp12 continuing RNA replication where it left off.
Exoribonuclease or no exoribonuclease
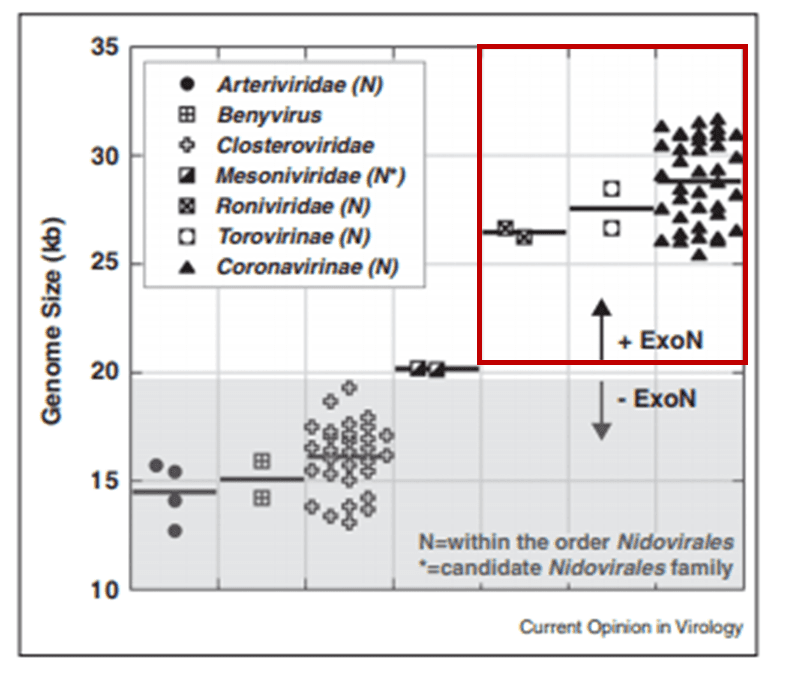
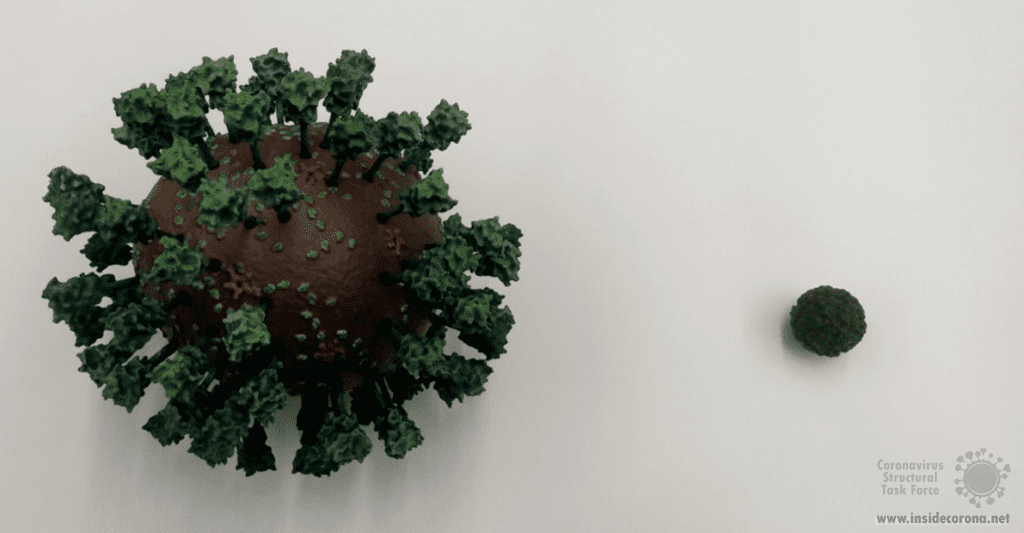
The proofreading ability from Helicase and nsp14 ExoN allows SARS-CoV-2 to have a huge genome as compared to other viruses6(Figure 2). The large 29.9 kb genome of SARS-CoV-2 requires much more physical space to accommodate the necessary genetic information for reproduction when compared to other RNA viruses, such as Rhinovirus that has a genome between 7.2 kb and 8.5 kb in size (Figure 3). When no ExoN proofreading is present genomes cannot expand beyond 20 kb in size6(Figure 2). Maybe by removing the exoribonuclease activity, irreversible damage could be caused to the genome of SARS-CoV-2.

Nsp14 Structure
In order to understand how nsp14 can do this, we need to find out its atomic structure; this may also allow us to develop a drug which hinders its function. However, to this date, no structure of nsp14 from SARS-CoV-2 has been solved. However, structures have been solved of nsp14 in complex with another viral protein, nsp10, both from SARS-CoV (PDB entries 5nfy, 5c8s, 5c8t, 5c8u)2,7. As the protein sequences are very similar between SARS-CoV and SARS-CoV-2 (nsp14 is 95%, and nsp10 is 97% identical), it can be assumed that the SARS-CoV-2 structure as well as its functionality are very similar to SARS-CoV. The active site of the ExoN domain of nsp14 from SARS-CoV-2 has a DEEDh motif (named for the one-letter codes of the amino acids involved) containing a histidine as well as two aspartates and two glutamates2,3,7,8.
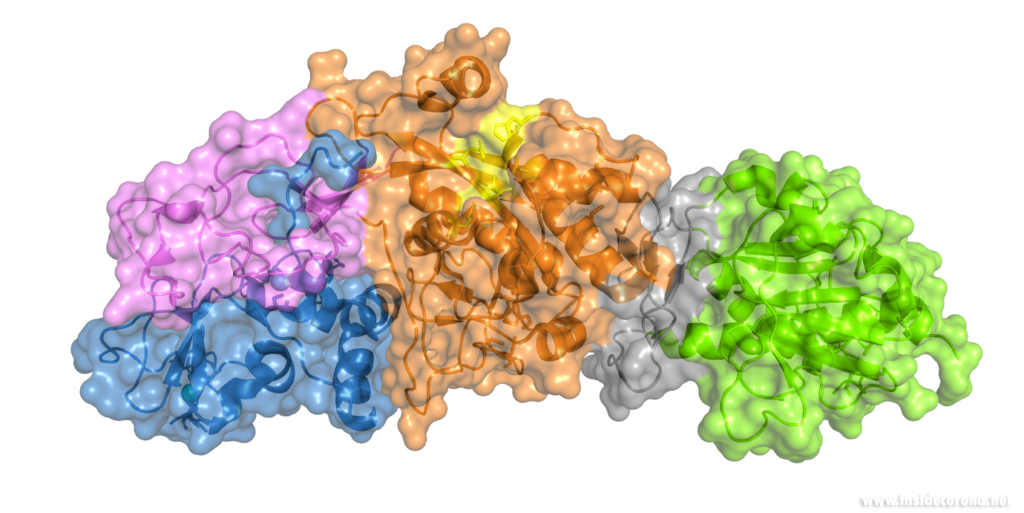
Nsp14 interacts with nsp10
The N-terminus of nsp14 interacts with nsp10 (pink and blue, respectively, in Figure 4). The following domain (orange) has been shown to have exoribonuclease activity on double stranded RNA in a 3’ to 5’ direction9. When nsp10 is interacting with nsp14 there is a 35 fold increase in exoribonuclease activity, which is thought to occur due to conformational changes caused by formation of the complex2,9. The ExoN domain of nsp14 (orange) is connected to the methyltransferase domain (green) by a flexible hinge (black)7,10. This flexible region opens up the methyltransferase active site to allow methylation of the N7 of the 5’ Guanosine triphosphate of RNA10. There are three zinc finger motifs in nsp14 with two found in the ExoN domain and one in the methyltransferase domain2,7. In combination with the two further zinc sites in nsp10, these zinc fingers hold loops of the proteins together and are involved with nucleotide interaction2,7.
Nsp14 has also been demonstrated to form complexes with the copy machinery , nsp12, nsp7, and nsp8, although this interaction is independent of nsp102,11,12.
Exoribonuclease active site and potential drug development
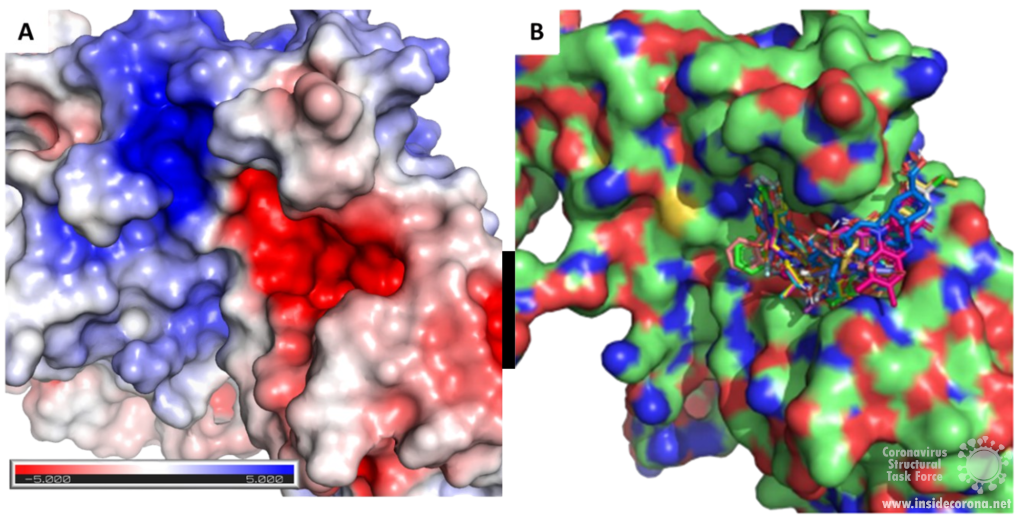
Scientists are searching for drugs that could be used to target nsp14 in order to find a cure for COVID-19. The active site of the ExoN domain of nsp14 has five residues that are essential for activity that form a negatively charged pocket (Figure 5A)7. Currently researchers are using the nsp14 structure from SARS-CoV to model a SARS-CoV-2 structure which can be used to identify compounds that could bind to the active site (Figure 5). These in silico screens start with nucleotide analog drugs like Remdesivir, Ribivarin or Ritonavir that are currently used as antiviral treatments for other viruses13–15. These nucleotide analogs are then changed to achieve a better binding to Nsp14’s active site in order to block it (Figure 5B).
As the ExoN is essential to support the huge 29.9kb genome of SARS-CoV-2, targeting nsp14 could lead to an effective treatment to COVID-19. Although drugs that target just nsp14 could be effective at increasing the error rate in RNA production by the virus, a more effective treatment will require inhibition of the RdRp of the copy machinery at the same time!
Available structures
If you would like to look at the currently available structures for Nsp14(currently only available from SARS-CoV), they are available from our data base; we provide information on the quality of measurement data and models as well as improved structures. The highest resolution structure of nsp14 is PDB entry 5c8t at 3.2Å. This has a bound S-Adenosyl methionine ligand as well as zinc atoms present. Alongside this, another structure of Nsp14 bound to S-Adenosyl homocysteine and a guanosine-triphosphate-adenosine ligand as well as zinc at 3.33Å resolution has been published (PDB: 5c8s). Additionally, two structures with zinc atoms but no ligands are available (PDB 5c8u 3.4Å at and 5nfy at 3.34Å). Both PDB entry 5c8t and 5nfy have improved structures re-refined by our group.
Sources
- 1.Zuo Y. Exoribonuclease superfamilies: structural analysis and phylogenetic distribution. Nucleic Acids Research. Published online March 1, 2001:1017-1026. doi:10.1093/nar/29.5.1017
- 2.Ferron F, Subissi L, Silveira De Morais AT, et al. Structural and molecular basis of mismatch correction and ribavirin excision from coronavirus RNA. Proc Natl Acad Sci USA. Published online December 26, 2017:E162-E171. doi:10.1073/pnas.1718806115
- 3.Barnes MH, Spacciapoli P, Li DH, Brown NC. The 3′–5′ exonuclease site of DNA polymerase III from Gram-positive bacteria: definition of a novel motif structure. Gene. Published online January 1995:45-50. doi:10.1016/0378-1119(95)00530-j
- 4.Chen Y, Cai H, Pan J, et al. Functional screen reveals SARS coronavirus nonstructural protein nsp14 as a novel cap N7 methyltransferase. Proceedings of the National Academy of Sciences. Published online February 10, 2009:3484-3489. doi:10.1073/pnas.0808790106
- 5.Chen J, Malone B, Llewellyn E, et al. Structural basis for helicase-polymerase coupling in the SARS-CoV-2 replication-transcription complex. Published online July 8, 2020. doi:10.1101/2020.07.08.194084
- 6.Smith EC, Denison MR. Implications of altered replication fidelity on the evolution and pathogenesis of coronaviruses. Current Opinion in Virology. Published online October 2012:519-524. doi:10.1016/j.coviro.2012.07.005
- 7.Ma Y, Wu L, Shaw N, et al. Structural basis and functional analysis of the SARS coronavirus nsp14–nsp10 complex. Proc Natl Acad Sci USA. Published online July 9, 2015:9436-9441. doi:10.1073/pnas.1508686112
- 8.Eckerle LD, Becker MM, Halpin RA, et al. Infidelity of SARS-CoV Nsp14-Exonuclease Mutant Virus Replication Is Revealed by Complete Genome Sequencing. Emerman M, ed. PLoS Pathog. Published online May 6, 2010:e1000896. doi:10.1371/journal.ppat.1000896
- 9.Bouvet M, Imbert I, Subissi L, Gluais L, Canard B, Decroly E. RNA 3’-end mismatch excision by the severe acute respiratory syndrome coronavirus nonstructural protein nsp10/nsp14 exoribonuclease complex. Proceedings of the National Academy of Sciences. Published online May 25, 2012:9372-9377. doi:10.1073/pnas.1201130109
- 10.Ogando NS, Ferron F, Decroly E, Canard B, Posthuma CC, Snijder EJ. The Curious Case of the Nidovirus Exoribonuclease: Its Role in RNA Synthesis and Replication Fidelity. Front Microbiol. Published online August 7, 2019. doi:10.3389/fmicb.2019.01813
- 11.Subissi L, Posthuma CC, Collet A, et al. One severe acute respiratory syndrome coronavirus protein complex integrates processive RNA polymerase and exonuclease activities. Proc Natl Acad Sci USA. Published online September 2, 2014:E3900-E3909. doi:10.1073/pnas.1323705111
- 12.Subissi L, Imbert I, Ferron F, et al. SARS-CoV ORF1b-encoded nonstructural proteins 12–16: Replicative enzymes as antiviral targets. Antiviral Research. Published online January 2014:122-130. doi:10.1016/j.antiviral.2013.11.006
- 13.Khater S, Dasgupta N, Das G. Combining SARS-CoV-2 proofreading exonuclease and RNA-dependent RNA polymerase inhibitors as a strategy to combat COVID-19: a high-throughput in silico screen. Published online June 24, 2020. doi:10.31219/osf.io/7x5ek
- 14.Shannon A, Le NT-T, Selisko B, et al. Remdesivir and SARS-CoV-2: Structural requirements at both nsp12 RdRp and nsp14 Exonuclease active-sites. Antiviral Research. Published online June 2020:104793. doi:10.1016/j.antiviral.2020.104793
- 15.Narayanan N, Nair DT. Ritonavir May Inhibit Exoribonuclease Activity of Nsp14 from the SARS-CoV-2 Virus and Potentiate the Activity of Chain Terminating Drugs. chemrxiv.org. Published May 13, 2020. https://chemrxiv.org/articles/Ritonavir_May_Inhibit_Exoribonuclease_Activity_of_Nsp14_from_the_SARS-CoV-2_Virus_and_Potentiate_the_Activity_of_Chain_Terminating_Drugs/12280043
Introduction
Have you heard that the coronavirus “mutates”? Or that there are “several strains” of it around the world? Sounds scary, right? However, the reality is that everything “mutates”. All organisms, over time, acquire differences in their genes, from bacteria to humans. You might be aware that this can happen when your DNA (Deoxyribonucleic Acid) is exposed to UV light (like from the sun!), but this can also happen during DNA replication. This is when a cell uses the template of one of the two DNA strands to make a new complimentary copy of the other strand. Mutation is common to all living organisms (and viruses) and a driver of evolution. This is the first post in a series that will explore coronavirus replication with a focus on the proteins involved.
How does the coronavirus make more of itself?
SARS-CoV-2 uses single-strand Ribonucleic acid (RNA) to encode its genome, not DNA, and hence belongs to a class of “single-strand RNA viruses”. For this reason, the virus needs a different way to copy its genome than “normal” cells have. The viral protein that copies the RNA is called an “RNA-dependent RNA polymerase” (RdRp). This protein uses the viral RNA as a template to make a new copy of viral RNA, by stringing single ribonucleotides together like beads on a string. This process is called polymerization.
A study by the Morse lab at Texas A&M University showed that SARS-CoV-2 RNA polymerase has a remarkable similarity to the RNA polymerase of SARS-CoV (>95%) as well as MERS-CoV [1], the virus which causes Middle-Eastern Respiratory Syndrome. This means that research performed in response to the SARS and MERS epidemics can inform our response to SARS-CoV-2. Unfortunately, a lack of consistent pandemic-preparedness funding means that we didn’t learn as much about RdRp in time as we could have. Still, RNA polymerase might be a viable drug target for halting the spread and reducing the fatality rate of COVID-19.
Structure of the RNA-Dependent RNA Polymerase
By determining the structure of RdRp, and deeply understanding how it works, we can optimize a drug to specifically target it and hinder its function. To this end, in the last few months, several structures of SARS-CoV-2 RNA polymerase have been published.
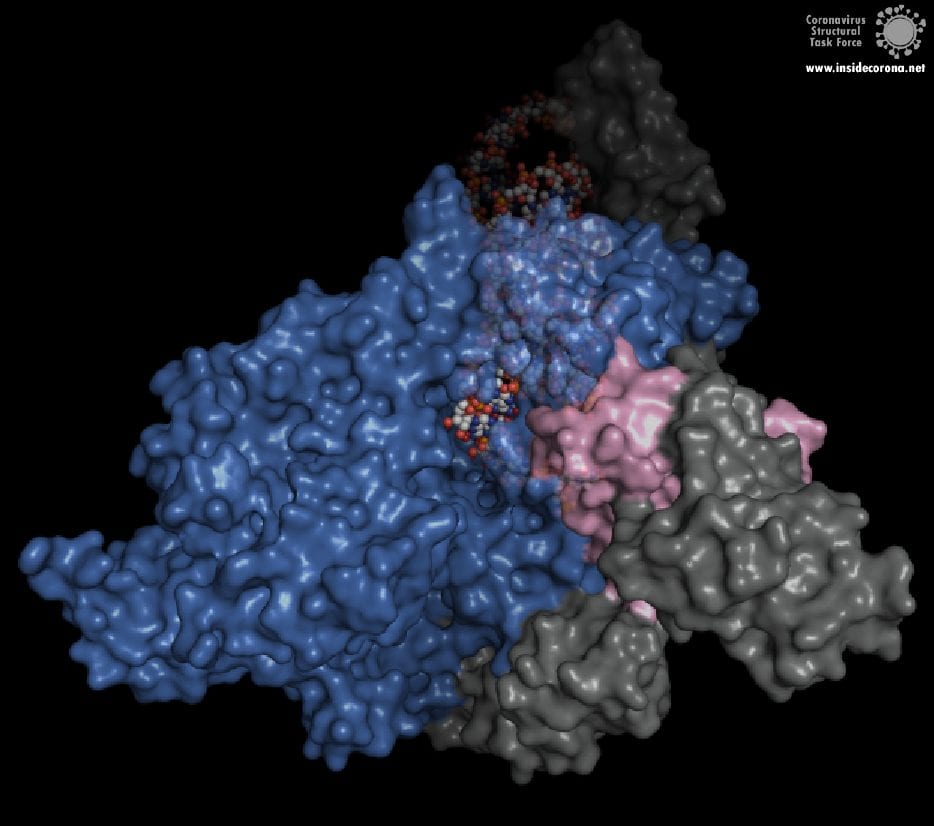
One interesting structure shows RNA polymerase in action, in the process of elongating an RNA strand (see Figure 1).[2] This structure clearly show the polymerase in complex with smaller proteins, non-structural protein 7 and 8 (nsp7 and nsp8). These proteins improve how well the RNA polymerase binds the template RNA and also how long it stays bound before dissociating – a feature called “processivity”.[3]
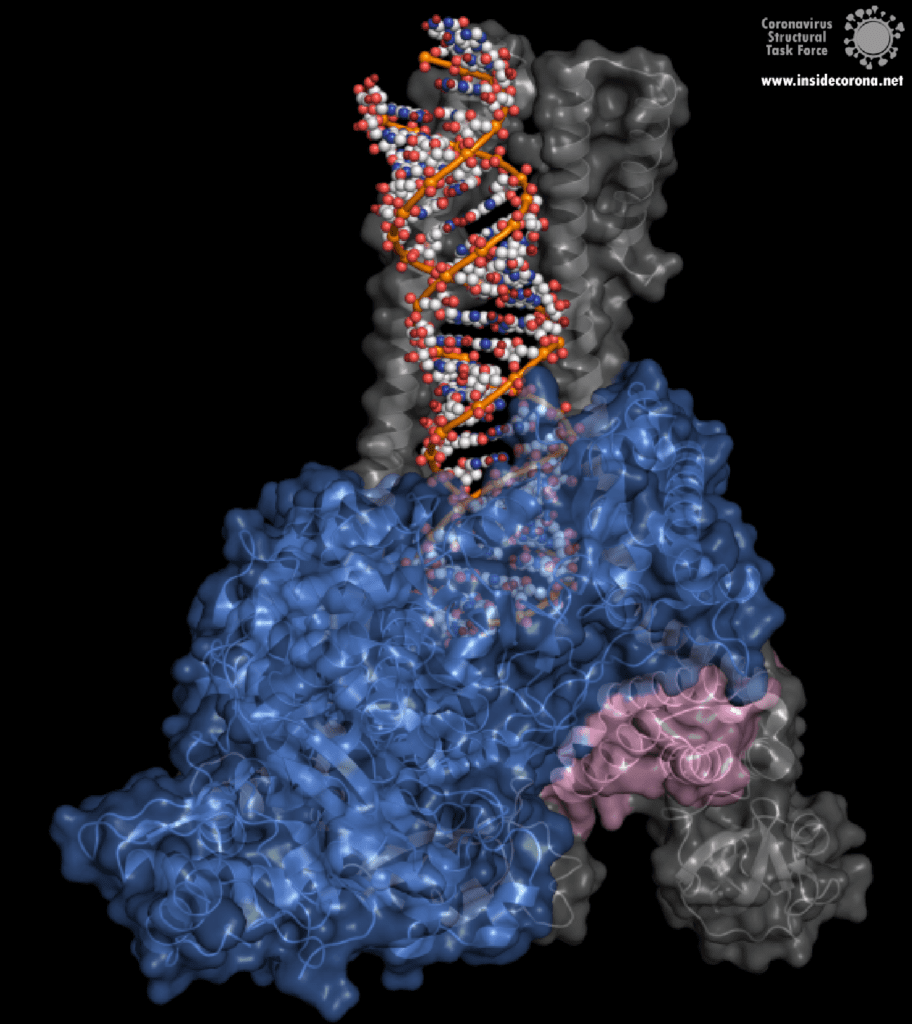
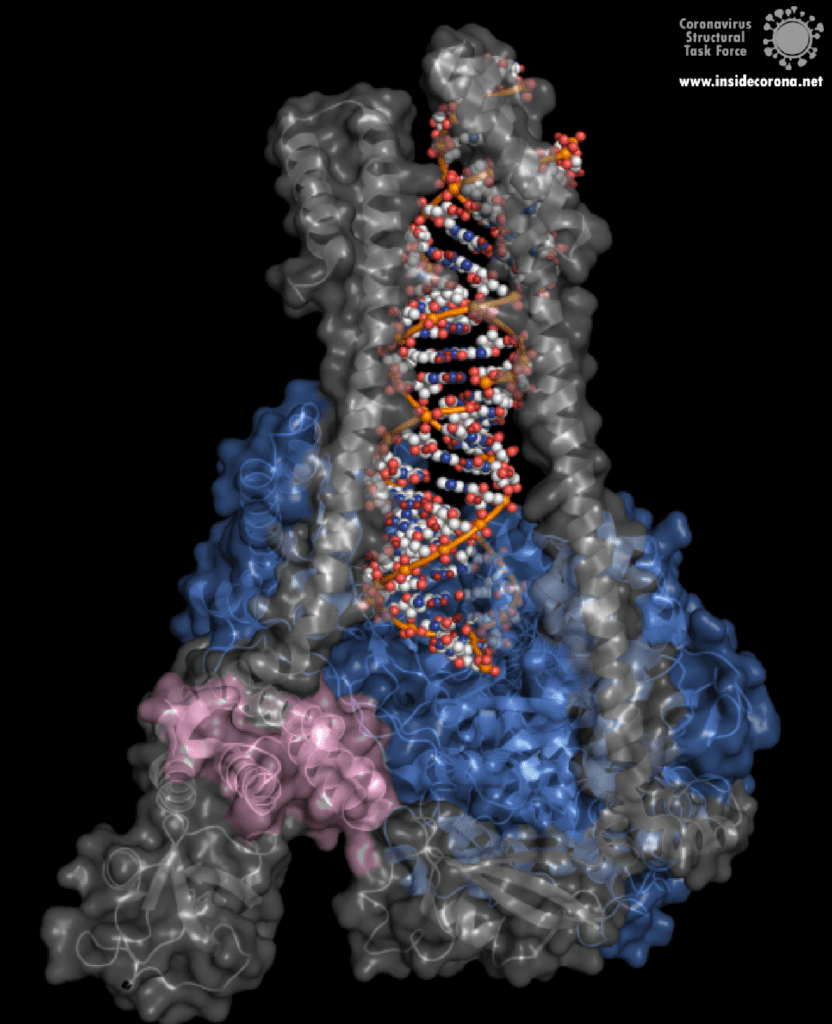
In the center of the protein is the area where the main action happens, called the “active site”. The amino acids of the polymerase that form the active site have a particular shape and chemical properties, which enable the polymerization reaction to occur very rapidly. In fact, the polymerase can string together as many as 100 nucleotides per second! [3] New RNA molecules can enter the active site through a little window to be added to the growing RNA chain. It is here that the antiviral drugs make their move!
How do antiviral drugs attack RNA-dependent RNA polymerase?
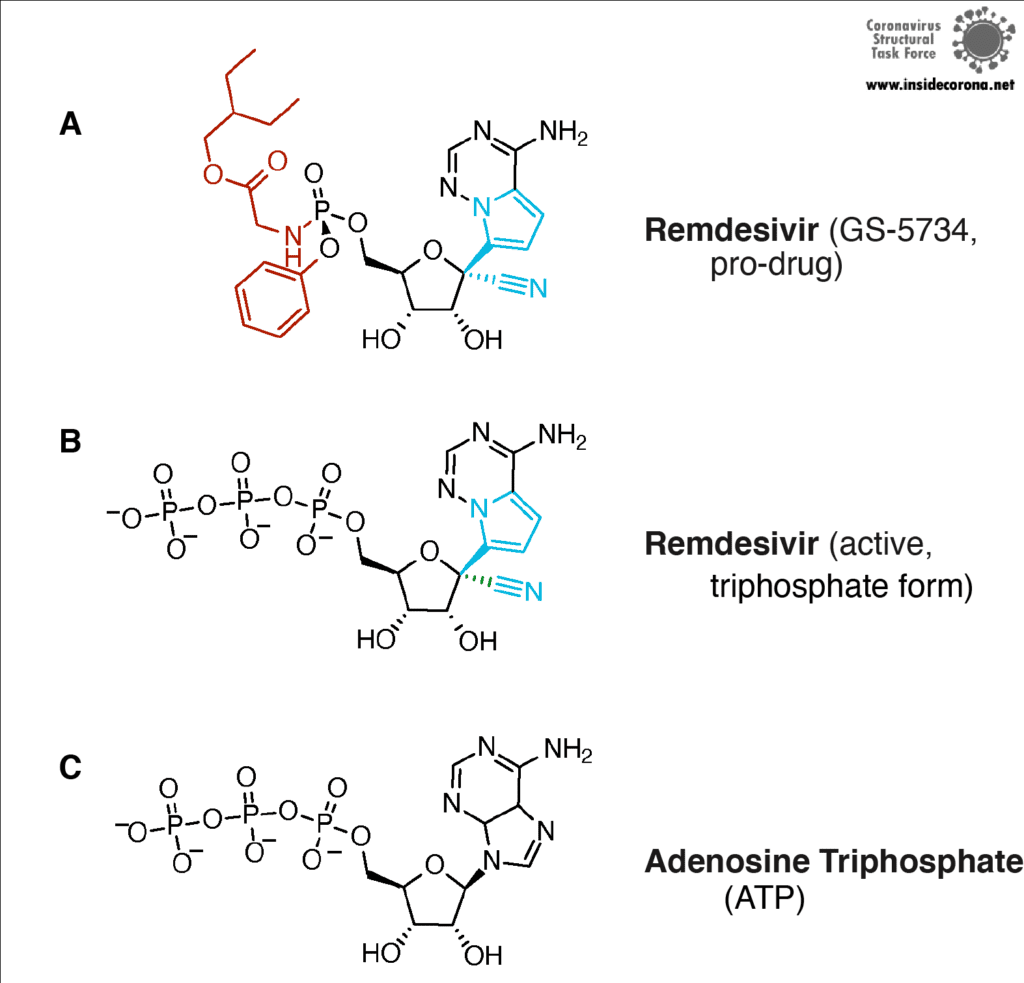
First, let’s talk about Gilead’s FDA-approved drug, Remdesivir, which has taken the spotlight in the search for COVID-19 cures. Remdesivir (which has a fancy chemistry ID, GS-5734, and is sold under the brand name Veklury), is a “nucleotide analog”, which means that it mimics the shape and chemistry of the nucleotides that make up RNA and DNA (see figure).
Remdesivir was developed originally as a general antiviral drug and was later shown to protect cells (in a test tube) and monkeys (not in a test tube) from the Ebola Virus [4]. However, this was recent enough, and science is slow enough that, until the COVID-19 pandemic, large-scale clinical trials of Remdesivir hadn’t been done yet. So scientists and doctors have been rushing to test the drug in COVID-19 patients. In fact, the US and Japan both approved the drug for “Emergency Use Authorization'' for severe COVID-19 patients as early as May [5], [6]. And, in July, the European Medicines Agency gave Remdesivir a “conditional marketing authorization” (used for drugs that meet an unmet medical need but have insufficient data for normal approval). This allows the use of Remdesivir in severe COVID-19 patients through the next year [7]. So, how the heck does a drug for Ebola, Influenza, or some other viruses also work against COVID-19? I was concerned by this when the news about all the drug trials were coming out – and I’m sure I wasn’t the only one...
The simple answer to that is all these viruses need to do the same thing - copy their RNA genome from an RNA template. And in order to do that, they all end up using basically the same tool, an RNA-Dependent RNA polymerase. And all drugs that are nucleotide analogs use the very same trick: they dress up like ribonucleotides (the "beads on a string" from before) and fool the RNA polymerase into letting them into the active site. Once inside, they get “stuck” in the active site, jamming the polymerase machine. Since this trick should work for any viral RNA polymerase, we can use these drugs for any RNA virus, and call them ‘general antivirals’. Of course, in practice, this doesn't always work, because there are differences between the different RNA polymerases. However, it is a great place to start! In the future, if we have general antivirals for SARS-CoV-2 all ready-to-go, we may be better equipped to deal with another coronavirus outbreak!

The Chemistry of Remdesivir
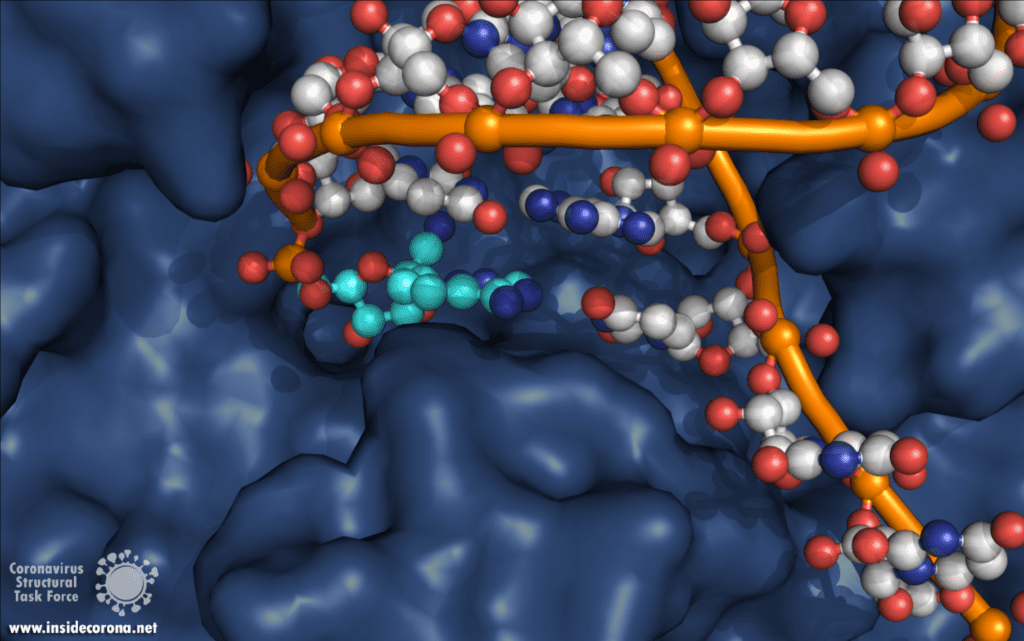
Remdesivir resembles the nucleotide adenine in structure, although it has some fancy chemical add-ons which help make it a better drug (thank you, medicinal chemistry!). When Remdesivir is injected into a vein, it travels through the bloodstream and enters into our cells, which recognize it as a foreign substance and try to digest it. However, what ends up happening is that the cells remove just the fancy chemical add-ons, and then confuse it for a normal adenine nucleotide. In infected cells, the viral RNA-dependent RNA polymerase then starts grabbing these molecules and inserting them into the new viral RNA strand in place of adenine molecules. Remdesivir, now attached to the RNA, jams the polymerase, rendering the virus unable to make more copies of its genome. Ultimately, this halts viral replication and helps the patient fight off the virus.
Another drug that inhibits the RNA polymerase activity is Favipiravir, sold under brand names Avigan, Abigan, and FabiFlu. Favipiravir has been discovered by Toyama Chemical Co., Ltd. in Japan and it has a similar mechanism to Remdesivir, except that it mimics a guanosine nucleoside instead of an adenine nucleotide [8]. This drug was approved in Japan back in 2014 for use in resistant cases of Influenza A and B, but still remains unapproved in the US (still in Phase II and Phase III clinical trials) and the UK [9]. This drug is also being tested for use against Ebola virus, Lassa virus, and currently SARS-CoV-2 in 43 countries. The approval of Favipiravir for COVID-19 has been much faster in China (Mar 15, 2020), Russia (Jun 3, 2020), and India (Jun 20, 2020)[10], [11]. Nonetheless, other countries, including Japan, are in various stages of clinical trials, and the results are anticipated to be out by the end of July [10].
So...do we have a cure for SARS-CoV-2?
Sadly, not yet. While the speed at which Remdesivir has gone through clinical trials is unprecedented, more work needs to be done to make sure it is safe and effective. Since (in the big scope of things) not a lot have people have taken Remdesivir, we aren’t really sure what all the side effects are, although there is emerging evidence for liver and kidney damage [12, 13]. The most common side effects are nausea (10% and 9% of patients), indigestion (7%) and increase of transaminases (6% and 8%). In one study, 3.6% of patients in a 10-day trial needed to stop taking therapy due to the latter. However, serious viral infections can also cause liver damage, so separating the two causes is a challenge! Remdesivir is not a cure-all, either. In one study it improved the recovery time from 15 days to 11 days, but it showed no effect for patients with mild to moderate disease, and no difference in median recovery time for patients who were already on a ventilator [14]. Since the drug has to be given by infusion over several days, there is a pretty small window in which Remdesivir can actually help.
Likewise, Favipiravir has its own side effects such as liver damage, elevated uric acid levels, kidney damage, skin allergies, etc. [15]. These effects restrict it for use by severe diabetes and heart patients. On top of that, it is not suitable for pregnant women because it can cause potential fetal deaths and deformities. It has been shown that Favipiravir works only during the earlier stages of SARS-CoV-2 infection when the body’s immune system isn’t totally drained, whereas it can result in a cytokine storm (when your immune system really freaks out) in severely ill patients. But, unfortunately, the virus doesn’t differentiate between humans while attacking, so a universal drug for COVID-19 has to be safe for use by all people.
However, these drugs are better than nothing, and by understanding the mechanisms involved, scientists can continue to improve upon the existing drugs for the benefit of all. While most of the ‘general antivirals’ that target RNA Polymerase have failed with SARS-CoV-2, Remdesivir has been relatively successful. Scientists think that this is actually because of a proofreading protein in SARS-CoV-2 called exonuclease. Immediately after the RNA-polymerase makes new RNA, exnuclease checks to make sure the new RNA is correct. In one study, another drug that mimics RNA called Ribivarin was shown to be removed from newly synthesized RNA by exonuclease [16]. Thankfully, Remdesivir is not excised , which is likely why it has been more successful than the other options [17], [18]. To read more about how nsp14 maintains the integrity and virulence of SARS-CoV-2, tune in to a future blog entry!

Recommended Structures
For those interested in reviewing the structures further, they are available in our GitHub repo, along with information about validation and, where relevant, improved structures. For a high-resolution comparison of the active site with and without Remdesivir, 7BV2 and 7BV1 (respectively) were published together at 2.5 and 2.8 Å. The elongating structure of the complex shown above (6YYT) has the polymerase as well as the cofactors and RNA very well resolved, with little "missing" density and a resolution of 2.9 Å. It is likely preferable to 6M71 and 7BTF, which were published with a similar resolution but with less of the complex resolved, and no RNA. For those interested, 7C2K and 7BZF (at 2.93 Å and 3.26 Å) show the complex bound to RNA in a pre- and post-translocation state.
Sources
[1] J. S. Morse, T. Lalonde, S. Xu, and W. R. Liu, “Learning from the Past: Possible Urgent Prevention and Treatment Options for Severe Acute Respiratory Infections Caused by 2019-nCoV,” ChemBioChem, vol. 21, no. 5, pp. 730–738, Mar. 2020, doi: 10.1002/cbic.202000047.
[2] H. S. Hillen, G. Kokic, L. Farnung, C. Dienemann, D. Tegunov, and P. Cramer, “Structure of replicating SARS-CoV-2 polymerase,” Nature, May 2020, doi: 10.1038/s41586-020-2368-8.
[3] W. Yin et al., “Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir,” Science, p. eabc1560, May 2020, doi: 10.1126/science.abc1560.
[4] R. T. Eastman et al., “Remdesivir: A Review of Its Discovery and Development Leading to Emergency Use Authorization for Treatment of COVID-19,” ACS Cent. Sci., May 2020, doi: 10.1021/acscentsci.0c00489.
[5] O. of the Commissioner, “Coronavirus (COVID-19) Update: FDA Issues Emergency Use Authorization for Potential COVID-19 Treatment,” FDA, May 04, 2020. https://www.fda.gov/news-events/press-announcements/coronavirus-covid-19-update-fda-issues-emergency-use-authorization-potential-covid-19-treatment (accessed Jul. 08, 2020).
[6] A. Sternlicht, “Japan Approves Remdesivir For Use On Severe COVID-19 Patients,” Forbes. https://www.forbes.com/sites/alexandrasternlicht/2020/05/07/japan-approves-remdesivir-for-use-on-severe-covid-19-patients/ (accessed Jul. 08, 2020).
[7] D. CZARSKA-THORLEY, “First COVID-19 treatment recommended for EU authorisation,” European Medicines Agency, Jun. 25, 2020. https://www.ema.europa.eu/en/news/first-covid-19-treatment-recommended-eu-authorisation (accessed Jul. 10, 2020).
[8] E. De Clercq, “New Nucleoside Analogues for the Treatment of Hemorrhagic Fever Virus Infections,” Chem. Asian J., vol. 14, no. 22, pp. 3962–3968, Nov. 2019, doi: 10.1002/asia.201900841.
[9] K. Shiraki and T. Daikoku, “Favipiravir, an anti-influenza drug against life-threatening RNA virus infections,” Pharmacol. Ther., vol. 209, p. 107512, May 2020, doi: 10.1016/j.pharmthera.2020.107512.
[10] T. Hornyak, “Japan sending Fujifilm’s flu drug favipiravir to over 40 countries for Covid-19 trials,” CNBC, May 04, 2020. https://www.cnbc.com/2020/05/04/fujifilms-flu-drug-favipiravir-sent-to-43-nations-for-covid-19-trials.html (accessed Jul. 14, 2020).
[11] G. P. Ltd, “Glenmark Becomes the First Pharmaceutical Company in India to Receive Regulatory Approval for Oral Antiviral Favipiravir, for the Treatment of Mild to Moderate COVID-19.” https://www.prnewswire.com/in/news-releases/glenmark-becomes-the-first-pharmaceutical-company-in-india-to-receive-regulatory-approval-for-oral-antiviral-favipiravir-for-the-treatment-of-mild-to-moderate-covid-19-855346546.html (accessed Jul. 14, 2020).
[12] Goldman, J. D. et al. Remdesivir for 5 or 10 Days in Patients with Severe Covid-19. N. Engl. J. Med. (2020) doi:10.1056/NEJMoa2015301
[13] Remdesivir Safety Forecast: Watch the Liver, Kidneys | MedPage Today. https://www.medpagetoday.com/infectiousdisease/covid19/86582
[14] J. H. Beigel et al., “Remdesivir for the Treatment of Covid-19 — Preliminary Report,” N. Engl. J. Med., vol. 0, no. 0, p. null, May 2020, doi: 10.1056/NEJMoa2007764.
[15] Sandhya Ramesh, “Favipiravir, Japanese drug that’s the new Covid treatment hope your chemist will soon stock,” ThePrint, Jun. 25, 2020. https://theprint.in/health/favipiravir-japanese-drug-thats-the-new-covid-treatment-hope-your-chemist-will-soon-stock/447987/ (accessed Jul. 14, 2020).
[16] F. Ferron et al., “Structural and molecular basis of mismatch correction and ribavirin excision from coronavirus RNA,” Proc. Natl. Acad. Sci., vol. 115, no. 2, pp. E162–E171, Jan. 2018, doi: 10.1073/pnas.1718806115.
[17] C. J. Gordon, E. P. Tchesnokov, J. Y. Feng, D. P. Porter, and M. Gotte, “The antiviral compound remdesivir potently inhibits RNA-dependent RNA polymerase from Middle East respiratory syndrome coronavirus,” J. Biol. Chem., Feb. 2020, doi: 10.1074/jbc.AC120.013056.
[18] L. Zhang et al., “Role of 1’-Ribose Cyano Substitution for Remdesivir to Effectively Inhibit both Nucleotide Addition and Proofreading in SARS-CoV-2 Viral RNA Replication,” bioRxiv, p. 2020.04.27.063859, Apr. 2020, doi: 10.1101/2020.04.27.063859.
