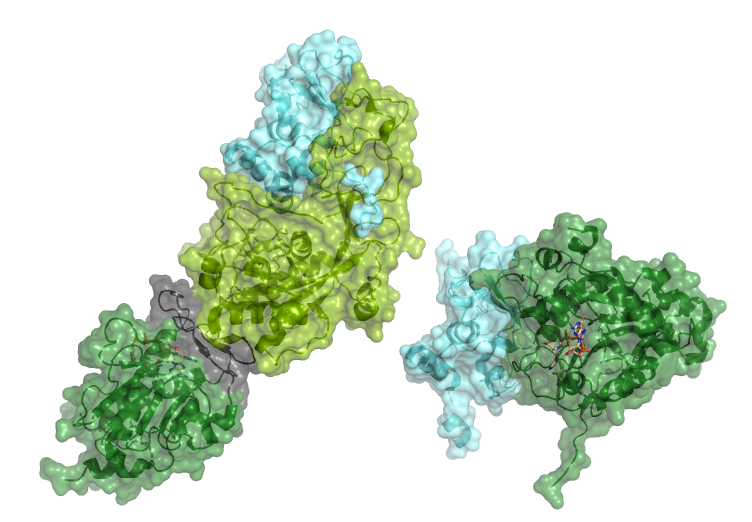
Two methyltransferase proteins, nsp14 and nsp16, have been identified within Coronavirus that perform methylation capping of RNA. These capping reactions are in important to help protect RNA from degradation, but also in mRNA processing and innate immunity. The protein nsp10 (light blue) is an activator for nsp14 and nsp16.
Nsp16 (dark green, right) has methyltransferase activity on the 2’-O of viral mRNA.
Nsp14 (also known as exonuclease) is a protein with dual function in viral replication and transcription.
There are two main domains present with a flexible linker (grey) connecting them. The N-terminal domain (light green) has an exoribonuclease function that proofreads the production of RNA in a 3’-5’ direction to prevent mutagenesis (see RNA polymerase complex). The presence of a proofreading enzyme facilitates the collosal genome of SARS-CoV2(29.8 kilobases in length).
Other RNA viruses such as Hepatitis C (9.6 kilobases) or Lassavirus (Two RNA strands, 3.4 kilobases each) do not encode an exoribonuclease enzyme due to their significantly smaller genomes.
The C-terminal domain (dark green) has methyl transferase activity for N7 capping of the 5’ end of viral mRNA. To date, no nsp14 from SARS-CoV2 has been solved however there are several structures from SARS-CoV which have 95% sequence identity to SARS-CoV-2 nsp14.