You don't want any more boring Christmas decorations? But something scientific? Back in 2021, we shortened our winter lockdown and decorated our windows with viral snowflakes.
This year, we've translated Dr. Hutchinson's craft template into German.
Since you are visiting our English site, you can just go ot the original source!
Dr. Hutchinsons Virus Snowflakes
Have fun cutting them out!
We wish you all a merry Christmas and a healthy 2023.

Knowing a protein’s 3D structure enables scientists to investigate its general shape and stability, deduce its potential function, and run drug-binding simulations to find a cure if the protein is from a pathogen. However, solving a structure is not an easy task, as proteins are too small to observe with optical microscopes. Furthermore, the process of collecting and preparing a protein sample and getting the final structure out of the experimental data can take several weeks or even months and is, hence, quite expensive.
In 2021, everything changed when AlphaFold2 was published: a new protein structure prediction software, which is not only incredibly accurate but also easy to use. Immediately, a great hype dominated the news, but did AlphaFold really change structural biology forever and will it make conventional methods obsolete?
Making Protein Structures Visible
Before we dive into the solution to an old problem of structural biology, we have to cover a few basics first. So, what are proteins again? In a nutshell, proteins are tiny nano-sized machines that perform any kind of task in your body. Some proteins organize the replication of your cells, while others digest your food. Another group of proteins makes up your hair, and plenty of other things in your body are also performed by proteins. A protein’s capabilities are determined by its shape, which raises great interest in its exact 3D structure. Unfortunately, proteins are smaller than visible light, so we cannot observe them with any optical microscope. Wait, smaller than light? How is this possible, you may ask? This picture should help to understand this:
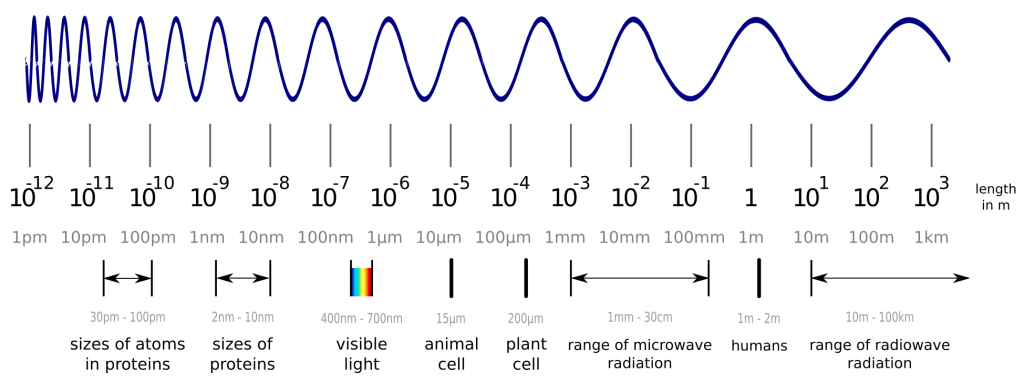
While visible light can easily interact with objects, which are larger than the wavelength of visible light itself, it does not interact much with smaller objects like proteins and atoms. Luckily, we can also create “light” in the invisible spectrum and measure the proteins with a detector instead of our eyes. One method to do this is X-ray crystallography. The details are complicated, but the important steps are to produce and purify your protein from bacteria, grow crystals out of your protein (yes, you can actually do this!) and shoot X-ray beams at it. With the data from the detector, you are able to create a 3D model of your protein, which may not represent reality perfectly, but is accurate enough to work with. You can find out more about these 3D models in another blogpost.
However, a lot can go wrong with this: The protein could kill the bacteria which should produce it for you, crystals might not form properly, and the process of turning the collected data into a model is not straightforward as well. All in all, this method can produce the desired result, but with a high investment of both, money and time.
The Protein Folding Problem
Proteins consist of long chains of amino acids. Most life on earth, us included, uses twenty different amino acids, each one with its own unique properties. For example, some are positively charged and like to be surrounded by negatively charged molecules or water. Others are not charged at all and prefer to hide inside a protein core and stay away from the water in which the protein is located. A protein’s amino acids are chained together in sequence and their properties define the shape, the final fold, of a protein. So, if you changed one amino acid in the chain into another one with vastly different properties, the whole protein would fold a bit differently. In fact, small changes might only modify the fold slightly, but multiple and huge differences in the sequence result usually in large changes in the fold. Since shape determines a protein’s function, the new protein might also do different things. The important takeaway here is: The information about the 3D structure is hidden in the sequence of amino acids. However, it remained a mystery how the folding into the final shape works in all details and could therefore not be replicated in simulations. This mystery is today known as the protein folding problem.
The hidden information in the amino acid sequence motivated many scientists around the globe to work on protein structure prediction software. Soon, those scientists started to compete against each other in the CASP competition – the Critical Assessment of Techniques for Protein Structure Prediction. Every two years since 1994, the state-of-the-art techniques have been measured against each other, but for decades all methods were not reliable enough and made many mistakes. Only recently, at CASP13 in 2018, Deepmind’s AlphaFold succeeded the first time with predictions of phenomenal quality for many of the given input sequences. Nevertheless, there was still much room for improvement. In 2020, AlphaFold2 followed and produced for the very first time in history structures almost as good as the conventionally solved ones. The protein folding problem itself has been known for more than 50 years now, but only with current technology and new approaches like the use of deep learning, structure prediction became finally real.
How accurate is AlphaFold?
When people talk about AlphaFold, they usually mean AlphaFold2. While scientists use calculations to measure the difference between a prediction and the experimentally solved structure it should resemble, the probably best way to demonstrate its accuracy to a newcomer in the field is a picture:
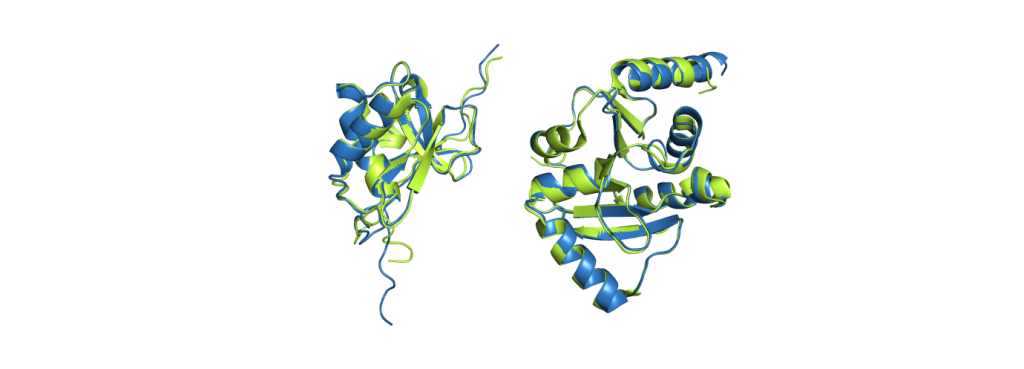
CoV-2 protein nsp3 (PDB code 7LGO), on the right is the Mac1 domain from the same protein (PDB code 6WEY).
The green structures are the ones obtained from experimental data; the blue structures are predicted by AlphaFold2. It is already a challenging task to predict which parts fold into helices and sheets (the spirals and the arrows), but AlphaFold2 even predicted a correct alignment of those and also the parts in between, resulting in nearly perfect agreement. The biggest deviations between experimental structure and prediction are usually found at the ends of the chains. While it does not give such excellent results for all proteins, it still performs pretty reliably and even provides some feedback on its confidence, making it easy to spot regions with a wrong fold.
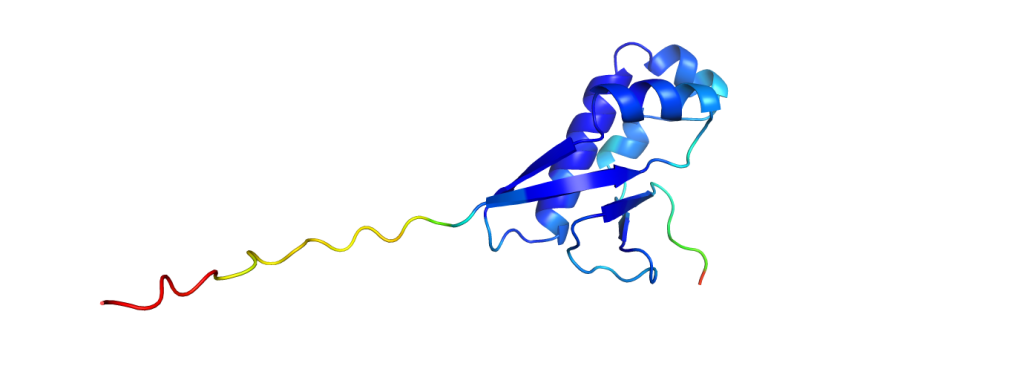
And to put the breakthrough of AlphaFold2 into perspective: to measure how similar the predictions are to experimentally solved structures, one could calculate a similarity value with one of many available metrics. One metric used in the CASP competition is the GDT, the global distance test, which returns values from 0% (no similarity at all) to 100% (identical structures). While all methods in the past did not pass on average the 60% mark, AlphaFold2 scored consistently with GDTs of over 90%.
Does AlphaFold replace conventional methods?
As we have seen, AlphaFold ist almost as accurate as the models obtained from X-ray crystallography. Does this mean that we no longer need those expensive and time-consuming methods? Well, for several reasons, this is not quite the case.
First and most importantly, predictions are not reality. They can help to simplify things or let us work into the right direction, but they cannot incorporate all the details of real biology. AlphaFold just takes a protein’s sequence of amino acids into account, but in reality, proteins are surrounded by water, small molecules and other proteins, and all of these affect the fold as well. Depending on the environment and on interaction with other molecules, some proteins even switch between multiple folds that are quite stable. Therefore, AlphaFold predictions are only a small part of the complete picture.
Another problem is posed by the so-called membrane proteins, a whole class of proteins which fix themselves at a membrane like, for example, the wall of a cell. Although AlphaFold does predict the individual parts across the membrane correctly, those are rarely aligned to each other correctly. This alignment problem occurs also with very huge proteins consisting of many smaller folded parts.
Last but not least, there are many proteins out there which were not predicted correctly by AlphaFold, so there’s still some room for improvement.
Important to mention is also the fact that AlphaFold was trained on the PDB, a database of experimentally solved protein structures. Without any new experimental data, prediction software like AlphaFold cannot be improved.
In summary, AlphaFold is good, but it is still not perfect.
In any case, the new predictions are useful not only to get a first glimpse of an unknown structure, but also to help solve structures by conventional methods. Remember that protein crystals are not always easy to grow? Well, this is sometimes due to regions which do not form a stable fold. AlphaFold2 can predict those regions and, thus, helps to design more successful experiments.
All in all, it is an incredible tool which does not only generate knowledge in a matter of hours instead of days but can also be used by everyone with no need for laboratory access or being an expert in structural biology. From here, we can only be curious what the next generation of structure prediction will be capable of, as AlphaFold2 already helped scientist around the globe to reveal the structural mysteries of various proteins.
The instructions and files below will allow you to create your own model of the virus! All you need is some spare time and a 3D printer. In addition, those without access to a 3D printer can still use the STL files to request printing from external services and then follow the instructions on assembling the same way. We do hope that this model will make the virus more tangible and that the model will not only be printed as a private project, but also be used for outreach activities and in educational institutions.
These instructions refer to the updated SARS-CoV-2 3D model released in early 2022 that considers new scientific insights and improves on the original. You can find details about the changes (and our reasons for them) in another blog post soon.
Our design is based on the best scientific evidence available. Not only are the shapes of the various proteins as close to the measured molecular structures as possible, but their numbers as well as the overall size of the virion match experimental results on a scale of 1:1,000,000. Therefore, 1 mm on the model represents 1 nm (10 Å).
By the way, this would make the RNA that is inside the virus hull 10 meters long and 1 mm thick.
What you need to get started
For easier printing and assembly, the virus structure has been broken down into individual components:
For the model with rigid spikes:
https://www.thingiverse.com/thing:5326932/files
- Virion (one Top and one Bottom)
The two virion components are completely solid irregular hemispheroids. The body has been broken down into two separate parts in order to produce two flat surfaces, minimizing the need for supports and lowering the amount of excess material when printing. The outer surfaces contain the necessary recesses for the spike proteins to sit in. This model contains 26 such holes. The surface of the virion is textured to represent SARS-CoV-2's E and M proteins.
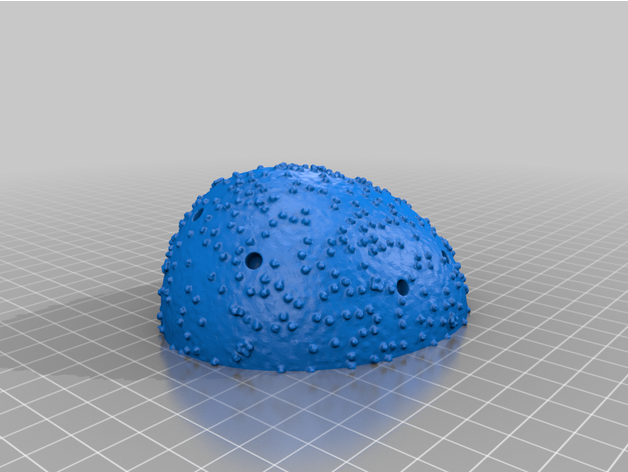
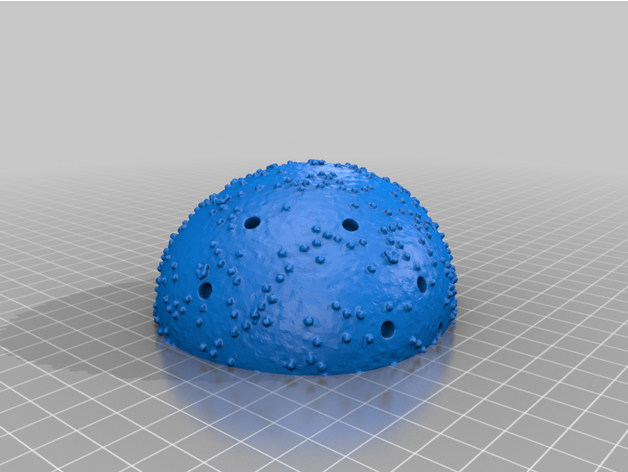


- 26 Spike Proteins
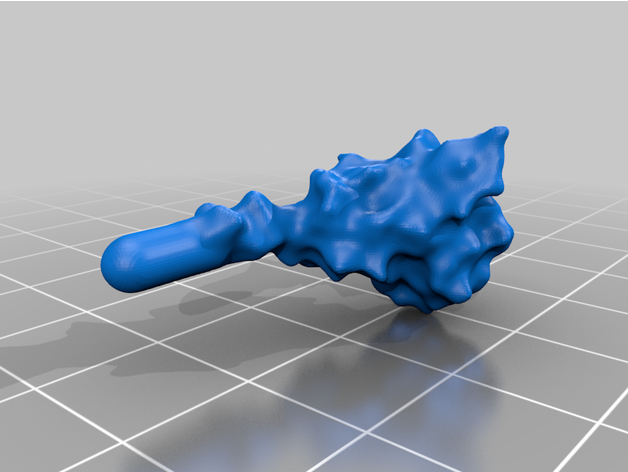
The spike protein stucture provides perhaps the most challenging aspect of the printing operation. Each spike consists of a complex crown-like surface and supporting stalk which connects it to the central virion. The individual spike STL files show the spike in different conformations and at varying angles.
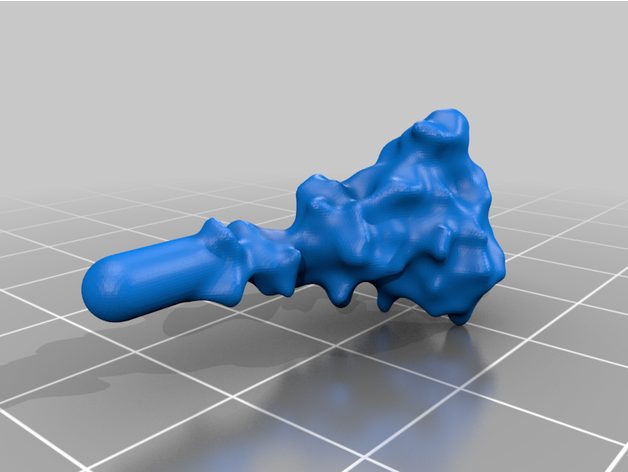
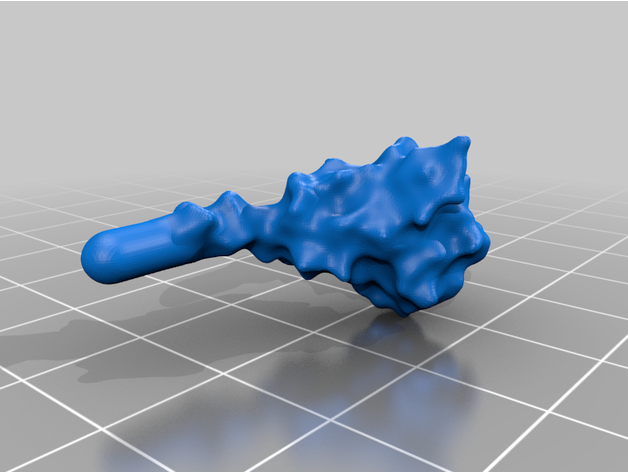
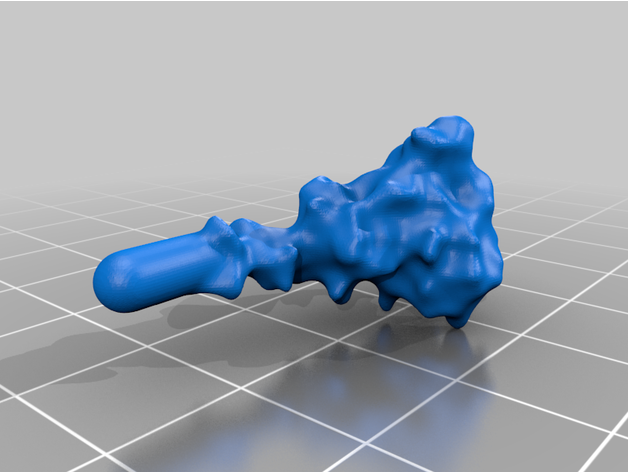
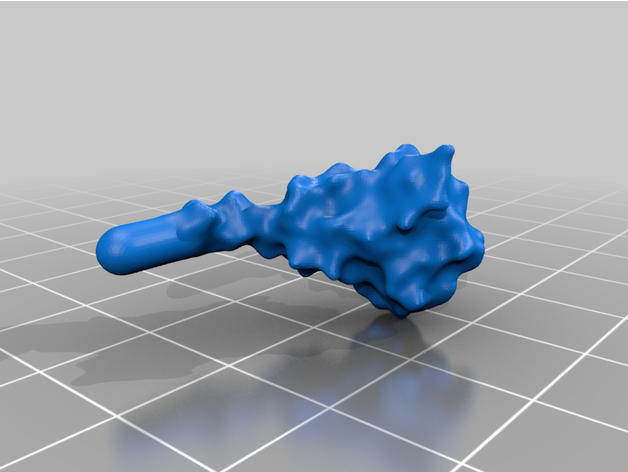
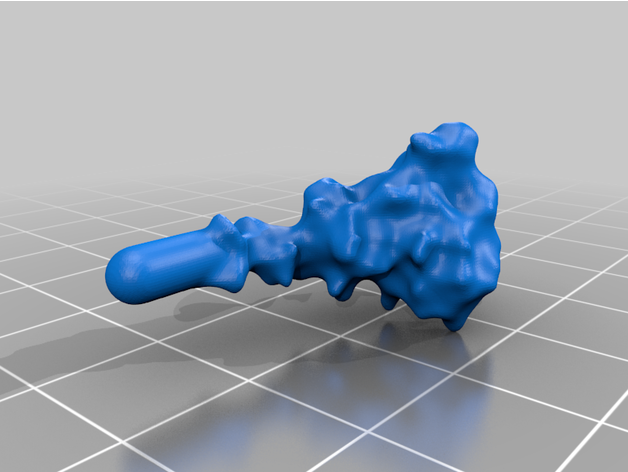






For the model with springs:
https://www.thingiverse.com/thing:5666273/files
- Virion (one Top and one Bottom)
The two virion components are completely solid irregular hemispheroids. The body has been broken down into two separate parts in order to produce two flat surfaces, minimizing the need for supports and lowering the amount of excess material when printing. The outer surfaces contain the necessary recesses for the spike proteins to sit in. This model contains 26 such holes. The surface of the virion is textured to represent SARS-CoV-2's E and M proteins.


- 26 Spike Proteins (11 extended, 15 retracted)
The spike protein stucture provides perhaps the most challenging aspect of the printing operation. Each spike consists of a complex crown-like surface that is mounted on a spring.
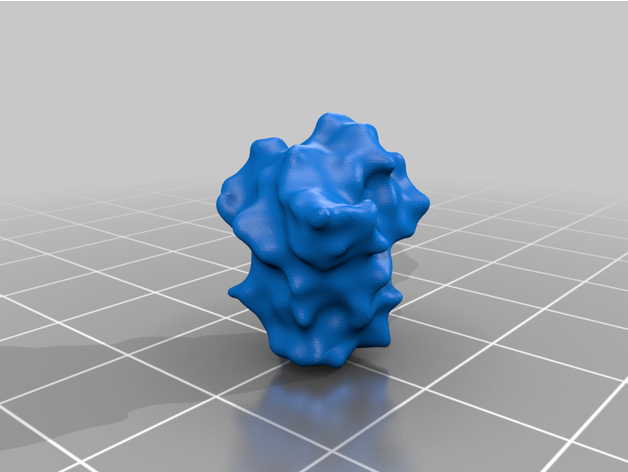
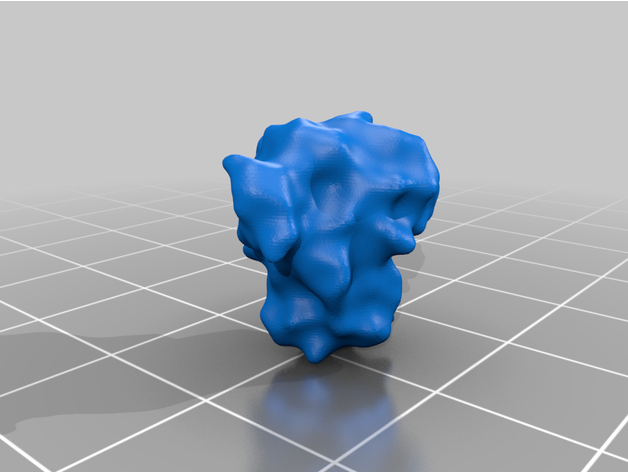


- Springs
You can purchase a suitable spring at https://www.mcmaster.com/compression-spring-stock/od~1-8/id~0-085/. This spring is 20 inches or 508 mm long. You can cut it either in 26 19mm pieces or 28 18mm pieces with two spare ones. If you want to get your springs from some other source, make sure that their outer diameter is about the same as this one (0.125 inches or 3.2 mm). The springs have to fit into the holes in the model parts.
To date, the structures have been printed successfully on several Fused Deposition Modelling (FDM) printers (Rostok MAX v2 & Prusa I3 MK3 printers), and we anticipate the even higher quality structures will be feasible with alternative methods, such as stereolithography (watch this space). Each of the parts is available in STL format and is printable through any suitable slicer software. Personal discretion is advised when setting up the prints, as the exact details may differ depending on conditions and equipment. The procedure outlined below will serve as a good starting point. Let us know of your experience in the comments!
Printing of the component parts
The first step is to print the individual components. For the body parts this is very straight forward as the surface negates the need for supports. The body objects can be printed with the minimum infill for support, though infill of 10% is recommended for rigidity.
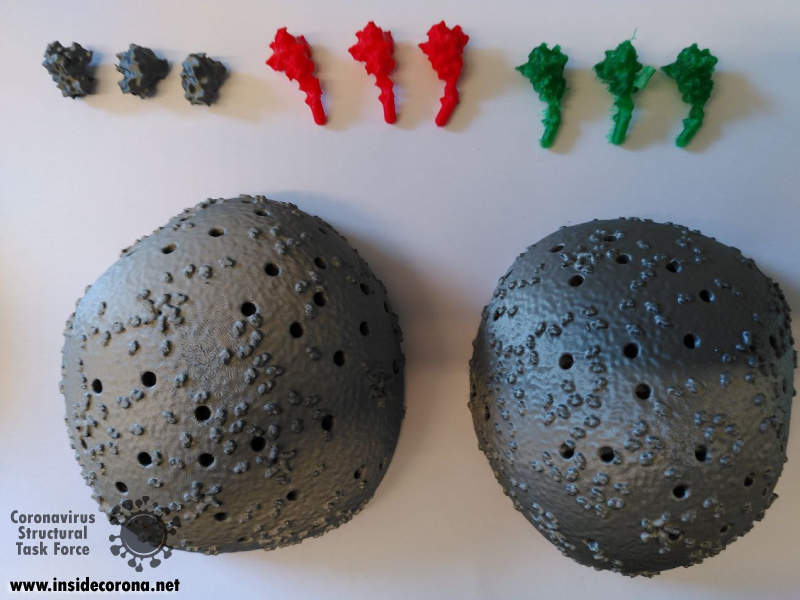

The spike proteins provide a more challenging print. The spike protein must be printed 26 times to complete the model. To represent the variety of conformations among the proteins in any given moment, we provide spikes in three different tilt angles in both the extended and the retracted state each. For a mixture of spike proteins that reflects a real-life virion reasonably well, we recommend this distribution:
For the model with rigid spikes:
- 3x 30° extended
- 4x 30° retracted
- 5x 40° extended
- 7x 40° retracted
- 3x 50° extended
- 4x 50° retracted
For the model with springs:
- 11x extended
- 15x retracted
The springs have to be about 3.2 mm in outer diameter and 18-19 mm in length. We recommend using stainless steel springs. To bend the springs, they are pulled onto several solid wires, which are bent into different spiral shapes. The wires with springs are then placed on a hot plate at 250 °C for 30 minutes. By using this method, different, random spring angles can be realized and the springs can be prevented from bending back into their original position.
We recommend printing the spike protein lying sideways as this results in stronger stalks. It is not too difficult to remove the supports of the stalks without breaking them.
We used FDM printing and ubiquitous poly-lactic acid (PLA), which made the post-processing easier.
A dual extruder printer would be ideal for spike printing as it would allow supports to be printed with water-soluble plastic, speeding up post-processing. In any case, printing individual or at least fewer spikes with greater spacing generally produces nicer objects which are easier to work with at the price of longer printing time.
Post-processing
Regardless of the approach taken for printing, some tidying will typically be needed to get the virion ready for assembly. Removing the supports can be done with a pair of pliers, while the smaller artifacts and issues will need brushing off or sanding. A dental pick can be quite useful.
For PLA, we found the best thing to clean and smooth the surfaces (after support removal) is ethyl acetate. While ethyl acetate is readily available in many chemical labs or at a pharmacy, acetone-free nail-polish remover offers a commercially accessible alternative. You should be using safety glasses and accurately fitting (!) gloves when handling ethyl acetate, ventilate the room well and, in case of skin contact, use a skin cream after washing your hands! It dissolves the plastic, breaking down the small extrusion artifacts on the surfaces and can be applied in many ways. We found it best to leave the parts in a sealed ethyl acetate vapor environment, e.g. by putting it in a smaller open vessel into a stainless-steel pot, which should be cleaned carefully afterwards. This technique results in even and clean results, though it will take up to a few days to fully smooth each object. The faster method is to simply submerge the small objects in ethyl acetate for 10-30 seconds, and then remove each object, leaving them to dry out on a surface. For the larger virion parts, the surface can be smoothed by rubbing it down with a cloth damped with ethyl acetate, which was also used to “weld” the two viral hull halves parts together. A small amount was dropped onto the flat surfaces on each section, before the two were pressed together until the plastic fused to become a single object. The seam was then smoothed down using the same process as before.
For acrylonitrile butadiene styrene (ABS), acetone may produce the same results.
Ready for Assembly!
Finally, the 3D model can be assembled. For assembling, the springs are first fixed with superglue in the holes of the body. UV resin is then used for their final fixation. The UV resin also serves as a filling material to completely close the holes. The spikes are attached to the springs in the same way as the springs are attached to the body. If the intention is to paint the model, we recommend assembling the model before starting with the coloring. In this way, the UV resin used to fill the holes can also be painted and a more visually appealing result can be achieved.
We hope that our adventure in 3D printing the coronavirus inspires you to give it a try! The process we described was completed in a little over a week. The printing jobs were completed in just over two days, the cleaning and post-processing took another two days, while the painting was done over the course of a weekend. This article provides a description of our technique and should provide enough detail on how, with the outlined necessary tools, you can create a similar result. The files have been distributed through Thingiverse under a Creative Commons BY-NC license: You may remix, adapt, and build upon this work non-commercially and acknowledge the "Coronavirus Structural Task Force" as original author.
https://www.thingiverse.com/thing:5326932
As with every 3D printed model, there are many different ways this could be tackled and achieved, and we look forward to seeing the many creative ways explored by others in this endeavor. Please do share experiences and results with us, either through the comments Thingiverse or on Twitter (you can tag us @thornlab).
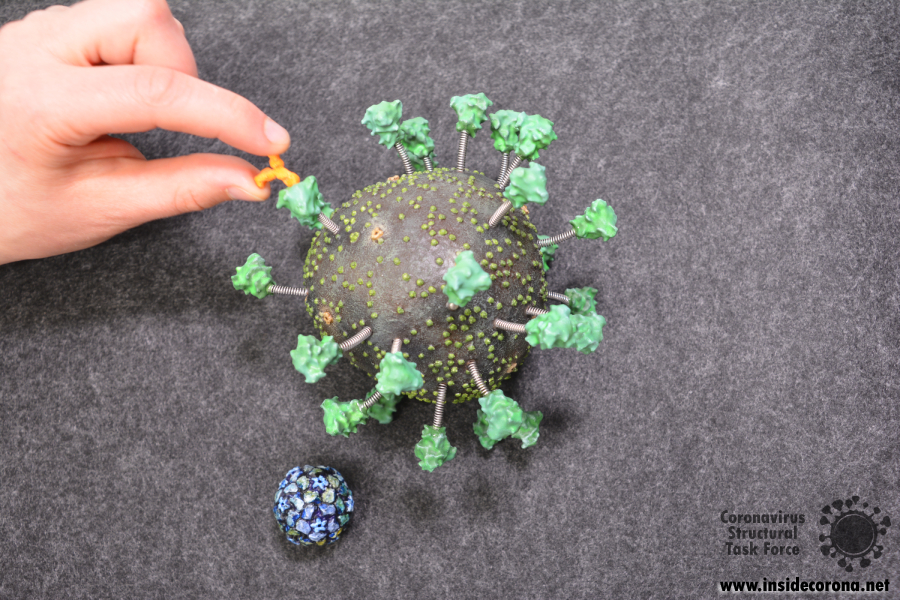

We have also designed a scale model of the human anti-body that binds to the spike protein. This is available alongside the virus model and can be attached to the spike protein as desired.
For a sense of perspective, we have also produced a model of the rhinovirus, which is one of the viruses that cause the common cold, at the same scale. It is available in STL format here: https://www.thingiverse.com/thing:4556845
Authors
We want to emphasize that the writing of this blog entry was a collaboration of a several people:
Dale Tronrud and Thomas Splettstößer worked together to create the STL files for the 3D model. Dale was the person to suggest it first (with Andrea Thorn picking up on the idea). Thomas then selected the experimental models and placed all the parts to form a realistic representation. Dale provided the knowledge about the limitations imposed by the nature of 3D printing and broke up Thomas’ model into printable parts that can be assembled without too much difficulty. He printed and assembled the first virion from this design. The updated model was printed at the facilities of the Physics department at the Universität Hamburg with generous support from PhysNET and Martin Stieben. Yunyun Gao and Philip Wehling refined the model, and Matthias Stäb painted the one shown in the pictures.
Our goal was to create a 3D printable model of SARS-CoV-2 that is as close to the actual virus as possible, but there are a lot of confusing and contradictory descriptions of the coronavirus SARS-CoV-2 in the literature. Combing through them in order to establish a complete understanding and a clear image of the virus that causes COVID-19 is a tremendous task and as new scientific observations come to light, our conceptions are being challenged, and need for adjustments arises.
At the start of the pandemic, when there were few measurements of the novel virus, its appearance was mainly inferred from knowledge about other coronaviruses, especially the closely related SARS-CoV-1 which caused the previous SARS pandemic [1]. The Coronavirus Structural Task Force created a printable model based on those images, but since then a lot of imaging data has been collected on the new virus, itself, and those images indicate that SARS-CoV-2 is sufficiently different. We have combed the literature and used what we learned to design an updated model.
The Shape of the Virus
Because viruses are so small, it is very difficult to directly observe them even with modern imaging techniques. (For more details, see here.) In the olden days, viruses were studied by coating samples with metal and then taking an electron microscopic image (see figure) so that the surface can be clearly seen. This is, by the way, the origin of the name “coronavirus”: The viral particles looked a bit like little suns surrounded by a corona of rays of light.
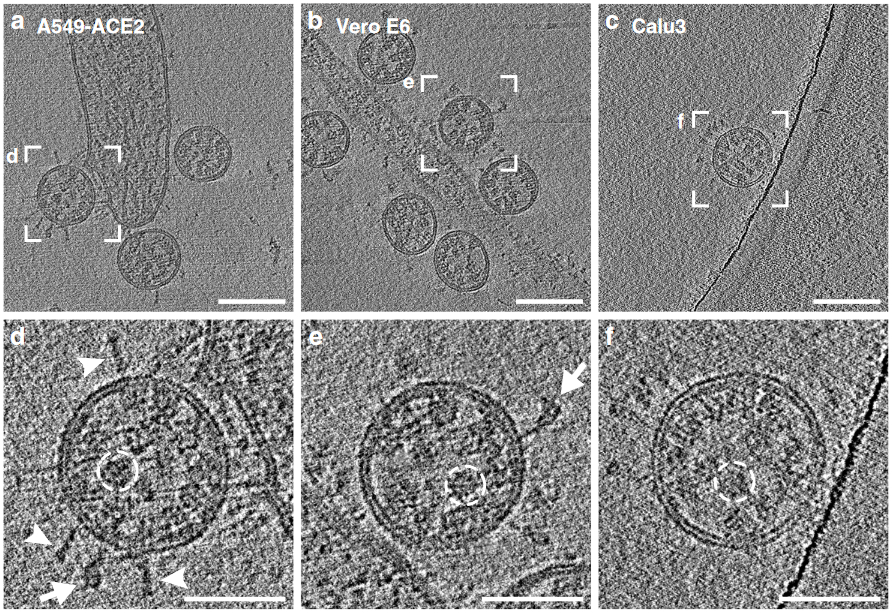
The most powerful ways to study the shape of a virus do not just investigate a single particle, but average the images of hundreds of them together resulting in information that does not describe a single virion but instead their average. In reality, and particularly for coronaviruses, every individual virus particle looks different, some being larger, and some smaller. They are only perfectly round in the absence of any exterior forces or perturbing internal structure. It does not take much to deform the “wobbly" thin double membrane hull of SARS-CoV-2. For example, it has been suggested that coronavirus particles are created in a close-to-perfect spherical shape and exposure to a slightly acidic environment causes virions to become a bit deformed, a change that might be important for infectiousness, and it has been postulated that the conformation of the M proteins affects the membrane curvature and, hence, the shape of the virus [2]. Our model is therefore not exactly round but more shaped like a potato. We do not claim that this is “the shape” of the virus, but simply one of many possible shapes.
A second difference between the old and new data is that SARS-CoV-2 appears to be smaller than the other viruses studied before this pandemic. To reflect the new data we have reduced the diameter of our model by 12%, from 100 mm to 88 mm. (Our model is scaled so 1 mm corresponds to 1 nm.) As with virual shape, individual viruses have a variety of diameters and 88 mm is simply one of many sizes which are consistent with the population. Due to the adjustments to the virus body its hull surface was reduced by roughly 23%. Research suggests that M proteins are distributed on the surface of a virion with a roughly constant density so we reduced their number by a matching 23%. The data for the E protein, however, indicates a roughly constant number in each virion so we have left their number at twenty. At this scale, the RNA contained in the virion would be about 10 meters long and one mm thick. One of the models for the virion body that we supply has a hole in the bottom that, both, allows the model to sit on a table and be used as a paper weight, and allows a 10 m long piece of twine to be hidden inside. The twine can be extracted during a demonstration to emphasize the surprising length of the SARS-CoV-2 RNA molecule.
Reconsidering the Spikes
As was found for the size of the virion, the experimental data derived from direct study of SARS-CoV-2 indicates a different number of spikes per particle than seen in previous studies, and the new number is far smaller. Current literature suggests the average number of spikes on the SARS-CoV-2 viral surface ranges from 26 [3] to 48 [4] . Our first model of SARS-CoV-2, based on images of other coronaviruses, had around 100 spikes but in our new model we have reduced their number to only 26.
While it is possible that SARS-CoV-2 does, indeed, have a different number of spikes than other coronaviruses, it is also possible that this substantial difference might well stem from changes in experimental approaches and circumstances. The microscopes and imaging techniques of today are much more powerful than those of even ten years ago.
In the past, despite the inconceivably small size of a single virion, coronavirus particles were too large to for all of its parts to be in focus in an image. Since one could only “see” the spikes in a limited region of a virus some rather elaborate methods were employed to estimate the total number of spikes. One of these is the Tammes problem, a mathematical puzzle that aims to distribute the maximum number of nonoverlapping, equal-sized circles on the surface of a sphere. Estimates for the minimal distance between two spike proteins on the viral surface are readily available even from 2D imaging, and following the assumption that the entire surface of the virion is covered with spikes packed as closely as possible the Tammes method suggests the presence of around 50 spikes per virus particle.

However, many images of SARS-CoV-2 show gaps between spikes, and it is quite likely that the density of spikes is lower. New techniques applied to SARS-CoV-2 can produce a sharp image of an entire, individual particle allowing the spikes to simply be counted. These studies show an average of 26 spikes per virion and this is probably the most reasonable estimate available until further experimental evidence emerges. This is just an average; the exact number of spikes on a particular virion varies around that value. The same publication also states that spikes are able to rotate freely on the viral surface and are not standing entirely upright, but are instead inclined by 40° on average [3], again varying a lot from spike to spike. Our updated version of the SARS-CoV-2 model incorporates these new findings. It has 26 spikes whose stalks are bent at angles of 30°, 40° and 50°. To improve the model’s flexibility and sturdiness, we also produced a version where the spike tops are connected to the body via springs. However, enthusiasts should remember that the spike proteins (as well as membrane and envelope proteins) are also “swimming” in the bilipidic membrane that serves as the outer hull, which we could not represent in our model.

Now, where’s the Model?
We made the updated files for the 3D-printable virus model available on thingiverse.
In this blog article you can find detailed construction guidance for the new model.
And here's a 3D preview of the new model on Sketchfab:
References
[1] "Coronavirus never before seen in humans is the cause of SARS". United Nations World Health Organization. 2003-04-16. Archived from the original on 2004-08-12. Retrieved 2022-02-17.
[2] Neuman, B., Kiss, G., Kunding, A. H. et al. A structural analysis of M protein in coronavirus assembly and morphology. J. Struct. Biol. 174, 1 (2011), 11–22. https://dx.doi.org/10.1016%2Fj.jsb.2010.11.021.
[3] Yao, H., Song, Y., Chen, Y. et al. Molecular Architecture of the SARS-CoV-2 Virus. Cell 183, 3 (2020), 730-738. https://doi.org/10.1016/j.cell.2020.09.018.
[4] Klein, S., Cortese, M., Winter, S.L. et al. SARS-CoV-2 structure and replication characterized by in situ cryo-electron tomography. Nat Commun 11, 5885 (2020). https://doi.org/10.1038/s41467-020-19619-7.
Why are vaccines developed so quickly and treatments so slowly?
In March 2020, the WHO declared COVID-19 a pandemic. Since then, 14 vaccines have entered the global market[1], and the number of immunized people grows every day. Even though vaccine development has never been this fast in history and will save many lives worldwide, people are still getting infected. To save patients afflicted with severe cases of COVID-19 and prevent health care systems from collapsing, effective drug treatments are the next step. But shouldn’t there already be drugs available?
What is the difference between vaccines and drug treatments?
While vaccines protect non-infected people from future infections, drugs help sick people. They alleviate symptoms and shorten the time to recuperate. Vaccines are important to combat a disease in the long run, through reaching herd immunity and protecting risk groups that would otherwise not survive an infection. Drugs, however, are equally important to fight acute infections.
How are vaccines developed?
First, scientists study the pathogen and its way of causing a disease in detail. Researchers then present a part of the virus that can be targeted by a vaccine for further investigation. This first lab stage is called the "research & discovery stage". What follows are pre-clinical trials, which are carried out in animals or cultured cell lines. Here, scientists prove that their vaccine candidate can cause an immune response and is not toxic.
If all data collected until this point looks promising, a clinical trial to test the vaccine in humans follows:
- Phase I examines the safety and dose for the vaccine, as well as the immune response.
- Phase II tests safety, efficacy, dosing, and immune response in a larger and more diverse group of people.
- Phase III tests the vaccine on thousands of subjects to determine efficacy and side effects.
- If successful, the vaccine is approved by the FDA in the US or the EMA in the EU. After approval, the vaccine is monitored in the real world, and more data is collected.
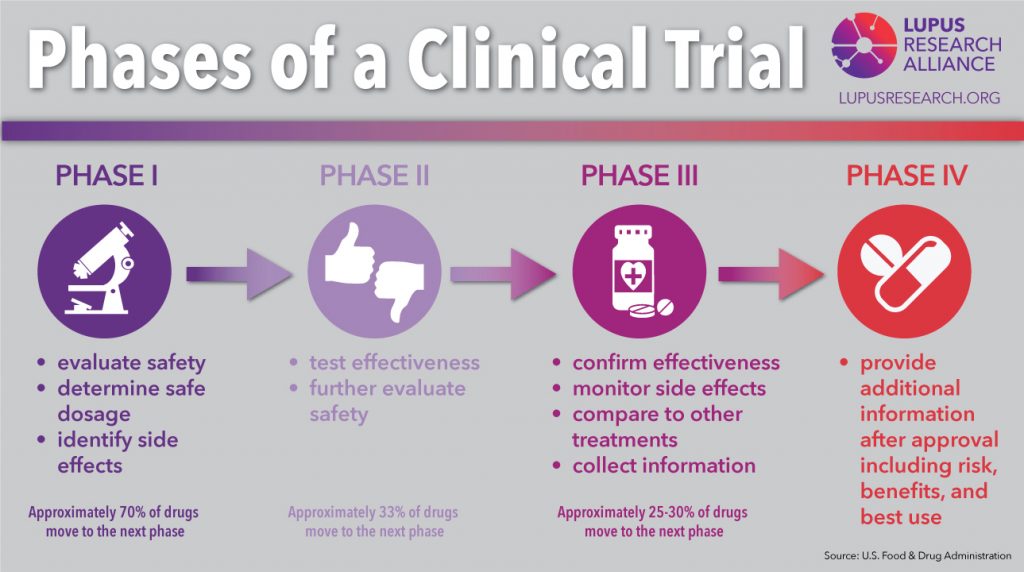
How come it progressed so fast for the coronavirus?
On January 10th, 2020, the genome of SARS-CoV-2 was published by a consortium of Chinese and Australian scientists[2]. This started global efforts to develop vaccines. For the first time, corporations, governments, and scientists from academia worked together on this scale to end the pandemic. The first vaccine approved in the western world was Pfizer-BioNTech in December 2020 (approved in the EU[3] and the US[4]). It took eleven months, a record time never reached for any other vaccine in history.
The fastest vaccine developed before COVID-19 was the mumps vaccine in the 1960s[5]. It took only four years to develop, which is amazingly fast compared to the average 10 years it takes for a vaccine to progress from basic research to approval[6]. Given this info, how was it possible to get SARS-CoV-2 vaccines ready in less than a year?
First, SARS-CoV-2 did not come out of nowhere, entirely. For years scientists have been studying its relatives SARS and MERS, their way of infecting cells, their proteins, and genetics[7]. Because of that, scientists did not have to start from scratch with researching these matters for the new coronavirus. Of course, there are differences, but the viruses’ general framework shows many similarities.
The cost of developing a vaccine, on average, surpasses the 1-billion-dollar mark[8], with a lot of the money spent on candidates that turn out to be failures. For many infectious diseases, funding simply cannot be sustained throughout development, especially if the disease is rare or occurs only locally. This was not a problem for the development of SARS-CoV-2 vaccines. Massive funding from governments and companies gave scientists more than enough resources to test their vaccine candidates. Also, because of all these resources, the developers were able to run several stages of testing in parallel[9].
Probably the most interesting reason for having vaccines this fast are the new vaccine technologies that have been developed since the turn of the century: mRNA and vector vaccines. Scientists often refer to them as vaccine platforms because they are technologies into which you only need to insert the part that is specific to a virus. For this, you need the virus’ genetic information, which was published early in this pandemic. Vaccine developers then inserted SARS-CoV-2 into their systems and were ale to quickly procede with the trials.
In the past, inactivated viruses or isolated viral proteins were used in vaccines. These, however, are highly specific to the virus you are trying to fight. This means, it was necessary to start at a basic level again every time a new vaccine was developed. Researchers hope that through these new vaccine platforms, the time for vaccine development in general will be a lot shorter in the future.
How is drug development different and why does it take so long?
The development of a new drug takes years, too. Generally, the timelines for drug and vaccine development show similar steps: Research & discovery, pre-clinical and clinical trials, approval and monitoring also apply for drug development.
Reaching the approval of an effective drug costs around $1,335.9 million[10], and failures are possible at every step of the way—like promising candidates from the pre-clinical stages showing no effect in humans, etc.
However, when comparing modern drug development to the new vaccine platforms, we can see just how much more complicated it is. Antiviral drugs are molecules that interact with parts of the virus and block it from entering a cell, stop it from replicating or stop another vital step of the infection path. There are also drugs that interact with parts of our own immune system to stop the disease from escalating.
Structurally, a drug candidate has to exactly fit its target, which is often a protein. This triggers two problems: What target should be chosen and what should the drug molecule look like.
Nowadays, millions of molecules are screened for their interaction with viral targets. This is done through computerized models but still takes a lot of time. Once there is a lead structure—a potentially effective molecule—, it is tested in pre-clinical trials and optimized structurally. During optimization, many hundreds of similar molecules are compared to see if their properties improve.
Different approaches to develop a drug take different amounts of time. One shortcut to developing an effective treatment is the repurposing of an already approved drug or a shelved candidate. The second fastest way is to develop a therapeutic antibody, followed by classic screening for a new drug.
Repurposing drugs for new diseases
Reusing drugs or drug candidates that have already been evaluated for safety greatly shortens the development time. It is even better when an already approved drug or a drug that is already backed with significant data from human trials shows an effect on the new disease.
An example for this is the first HIV medication AZT. It was first developed in 1964 as a potential cancer therapy, but later was found not to be very effective. In the 1980s, it was included in a screening for AIDS treatment and was found to interfere with HIV’s replication. It was later shown to decrease the death rate in people with AIDS and subsequently approved for treatment[11].
Remdesivir—a dead end?
Remdesivir is an antiviral drug that was designed to interfere with the replication of the genome of RNA-based viruses. The drug was first developed as a potential treatment for hepatitis C and respiratory syncytial virus and was later tested against the Ebola virus[12], which did not lead to convincing results[13].
But what do COVID-19 studies with remdesivir say?
The EU and US approved this medication in 2020 based on trials with patients that had moderate or severe cases. One study found that hospitalized patients with moderate COVID-19 benefited from a 5-day treatment with remdesivir.[14] For severe cases, there is some evidence that remdesivir could shorten the time to recuperate better than a placebo[15].
But the WHO is now advising against using the drug based on a meta-analysis they did. This convinced the EMA to re-evaluate remdesivir and maybe even take back the approval in the EU[16]. In the WHO study, the effect of four repurposed COVID-19 treatments—including remdesivir—on 11,330 adults was examined[17]. In this analysis, remdesivir showed little to no effect on hospitalized patients with COVID-19, as indicated by overall mortality, need of artificial ventilation, and length of hospital stay.
The other repurposed drugs from the WHO analysis were hydroxychloroquine (a treatment used against malaria), Lopinavir (a protease inhibitor used against HIV), and interferon beta-1a (an immune modulating drug for MS treatment). None of these showed a beneficial effect against acute COVID-19.
Dexamethasone is a success
The most dangerous aspect of a COVID-19 infection is that the person’s immune system overreacts and attacks the body in addition to the virus. Steroids are often used to dampen an immune response, so the commonly available drug dexamethasone was tried as a treatment. This has proven to be highly effective and has now become part of the standard care for COVID-19 patients[18].
Other drugs currently under evaluation
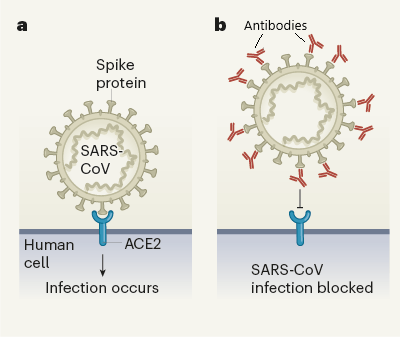
Monoclonal antibody treatments are currently under review in the EU and US.[19] Their effect: When the antibody attaches to the spike protein, the virus cannot enter the body’s cells, depriving it of its means of replication. Some of these treatments are combinations of two different antibodies that can attach to different parts of the spike, in theory, binding it more effectively.
The three treatments currently in the EMA review pipeline are: bamlanivimab and etesevimab, REGN-COV2[20], and regdanvimab[21]. The FDA[22] has authorized several monoclonal antibodies for emergency use and they have been shown to be effective at reducing the symptoms of COVID-19 if administrated early in the course of the disease[23].
Ongoing efforts
As there is more and more structural data available on SARS-CoV-2’s proteins, there are also more interaction studies with potential drug molecules and combinations.
Still, drug development takes time. In comparison to vaccines, which have just had a technology revolution, drugs will take longer from basic research to being ready for use. But there are already many different types of drugs under development, since funding is not a problem for COVID-19 treatments, and international collaboration speeds up the process as well.
Whether any of the current candidates will prove effective in treating COVID-19, remains to be seen. Maybe even a combined therapy with multiple drugs could be used to achieve the desired outcome.
[1] https://www.unicef.org/supply/covid-19-vaccine-market-dashboard
[2] https://virological.org/t/novel-2019-coronavirus-genome/319
[3] https://www.ema.europa.eu/en/medicines/human/EPAR/comirnaty
[4] https://www.fda.gov/emergency-preparedness-and-response/coronavirus-disease-2019-covid-19/pfizer-biontech-covid-19-vaccine
[5] Tulchinsky, Theodore H.. “Maurice Hilleman: Creator of Vaccines That Changed the World.” Case Studies in Public Health (2018): 443–470. doi:10.1016/B978-0-12-804571-8.00003-2
[6]COVID-19 vaccine development pipeline gears up. Mullard, Asher. The Lancet, Volume 395, Issue 10239, 1751 - 1752
[7] Abdelrahman Zeinab, Li Mengyuan, Wang Xiaosheng. Comparative Review of SARS-CoV-2, SARS-CoV, MERS-CoV, and Influenza A Respiratory Viruses. Frontiers in Immunology 11 (2020). doi:10.3389/fimmu.2020.552909
[8]Estimating the cost of vaccine development against epidemic infectious diseases: a cost minimisation study, Gouglas, Dimitrios et al. The Lancet Global Health, Volume 6, Issue 12, e1386 - e1396
[9] https://www.nature.com/articles/d41586-020-03626-1
[10] Wouters OJ, McKee M, Luyten J. Estimated Research and Development Investment Needed to Bring a New Medicine to Market, 2009-2018. JAMA. 2020;323(9):844–853. doi:10.1001/jama.2020.1166
[11] https://www.niaid.nih.gov/diseases-conditions/antiretroviral-drug-development
[12] https://www.gilead.com/-/media/gilead-corporate/files/pdfs/covid-19/gilead_rdv-development-fact-sheet-2020.pdf
[13] Pardo, Joe et al. “The journey of remdesivir: from Ebola to COVID-19.” Drugs in context vol. 9 2020-4-14. 22 May. 2020, doi:10.7573/dic.2020-4-14
[14] https://jamanetwork.com/journals/jama/fullarticle/2769871
[15] https://www.nejm.org/doi/10.1056/NEJMoa2007764
[16] https://www.ema.europa.eu/en/news/update-remdesivir-ema-will-evaluate-new-data-solidarity-trial
[17] Repurposed Antiviral Drugs for Covid-19 — Interim WHO Solidarity Trial Results. 384, 497-511 (2020).
[18] https://www.covid19treatmentguidelines.nih.gov/immunomodulators/corticosteroids/
[19] https://www.ema.europa.eu/en/human-regulatory/overview/public-health-threats/coronavirus-disease-covid-19/treatments-vaccines/treatments-covid-19/covid-19-treatments-under-evaluation
[20] https://www.ema.europa.eu/en/news/ema-starts-rolling-review-regn-cov2-antibody-combination-casirivimab-imdevimab
[21] https://www.ema.europa.eu/en/news/ema-starts-rolling-review-celltrion-antibody-regdanvimab-covid-19
[22] https://www.fda.gov/consumers/consumer-updates/know-your-treatment-options-covid-19
[23] https://www.fda.gov/news-events/press-announcements/coronavirus-covid-19-update-fda-authorizes-monoclonal-antibodies-treatment-covid-19-
Since the outbreak of SARS-CoV-2, infection has continued to spread. At the same time, governmental agencies around the world have adjusted the rules to prevent its spread. Information sources as basis for these rules have been obtained from scientific studies, public health research and simulation tests to understand the efficiency of mask types in preventing spread of infection by SARS-CoV-2. In this article, we will look at the mask types in use today, how much they can impede viral droplets and aerosols and how the construction of different masks helps to protect us from infection by SARS-CoV-2.
SARS-CoV-2 droplet sizes and viral transmission
The SARS-CoV-2 virus can be transmitted via droplets and aerosols.
Droplets are particles of sizes varying from 0.05 to 500 μm. They are directly emitted while breathing or talking. After being released into the air, larger droplets fall to the ground and others rapidly evaporate to form droplet nuclei less than 5 µm of size, also called aerosols, containing viruses in the range of 0.02 to 0.3 μm. Droplet nuclei can remain suspended in air for a longer time compared to large droplets and potentially contribute to airborne transmission1,2,3.
SARS-CoV-2 has been observed to be transmitted via 3 modes:4,5,6
- Contact transmission (usually via direct contact with infected persons, surfaces, or air)
- Droplet transmission over short distances when a person is close to an infected person
- Aerosol transmission over longer distances via inhalation of aerosols that remain airborne and travel with the air
Although maintaining a safe distance from an infected or possibly infected person will prevent viral spread via direct contact and droplet transmission, maintaining a safe distance may not be able to prevent spread of infection through airborne aerosols. This is why it becomes even more important to wear a mask.
Mask types and structure
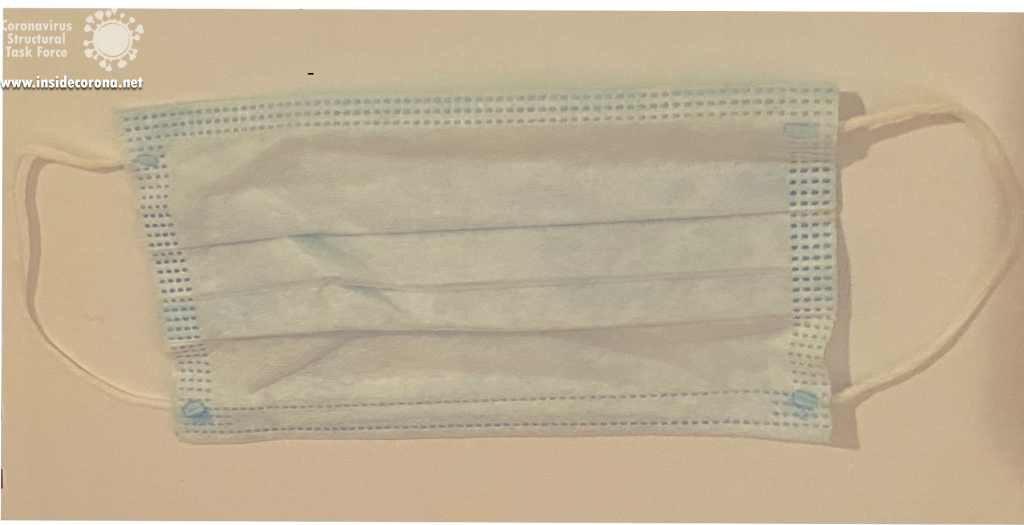
Surgical masks, also called medical face masks or mouth-nose protection (MNS), are disposable products that are normally used in clinics or in doctor's offices on a daily basis. They are made of special plastics with multiple layers. They have a rectangular shape with wrinkles so that the mask can adapt to the face. The front (outside) is often coloured, the back (inside) is not. The masks have ear loops and a wire noseband (see Figure 1).
Due to the shape and fit of most medical face masks, some of the breathing air can flow past the edges. Especially during inhalation, unfiltered breathing air can be sucked in. Therefore, medical face masks usually offer the wearer less protection against pathogenic aerosols than particle-filtering half-masks (FFP). Medical face masks, however, can protect the mouth and nose of the wearer from pathogen transmission via direct contact, for example with contaminated hands.
Since they are medical devices, their manufacturing and distribution must be carried out in accordance with medical device law. They must therefore comply with the legal requirements and the European standard EN 14683:2019-10. Only then can manufacturers mark the medical masks with the CE mark and distribute them freely in Europe. This is subject to supervision by competent authorities7.
Particle filtering half masks / filtering facepieces (FFP) are objects of personal protective equipment (PPE) within the framework of occupational health and safety. They protect the wearer of the mask from particles, droplets, and aerosols. When worn correctly, FFP masks are tightly attached and offer external and self-protection. Since the masks are disposable products as intended by the manufacturer, they should be changed regularly and disposed of after use.
FFP masks are produced either with or without an exhalation valve. Masks without exhalation valve filter both the inhaled air and the exhaled air over the mask surface and therefore offer both self-protection and external protection. Masks with valves offer less external protection because exhaled aerosols are not intercepted by the filter material but are only slowed down and swirled to a certain extent by the valve.
Like medical face masks, FFP masks must comply with clear requirements of laws and technical standards. In particular, the filter performance of the mask material is tested with aerosols in accordance with the European standard EN 149:2001+A1:2009. FFP2 masks must filter at least 94% of the test aerosols, for FFP3 masks the minimum is even 99% . They are therefore proven to provide effective protection against aerosols. The test standard, together with the CE mark and the four-digit identification number of the notified body, is printed on the surface of the FFP mask7.

Mask standards
The table below shows the currently accepted standards for masks and how they are effective in filtering out bacteria as well as particles.
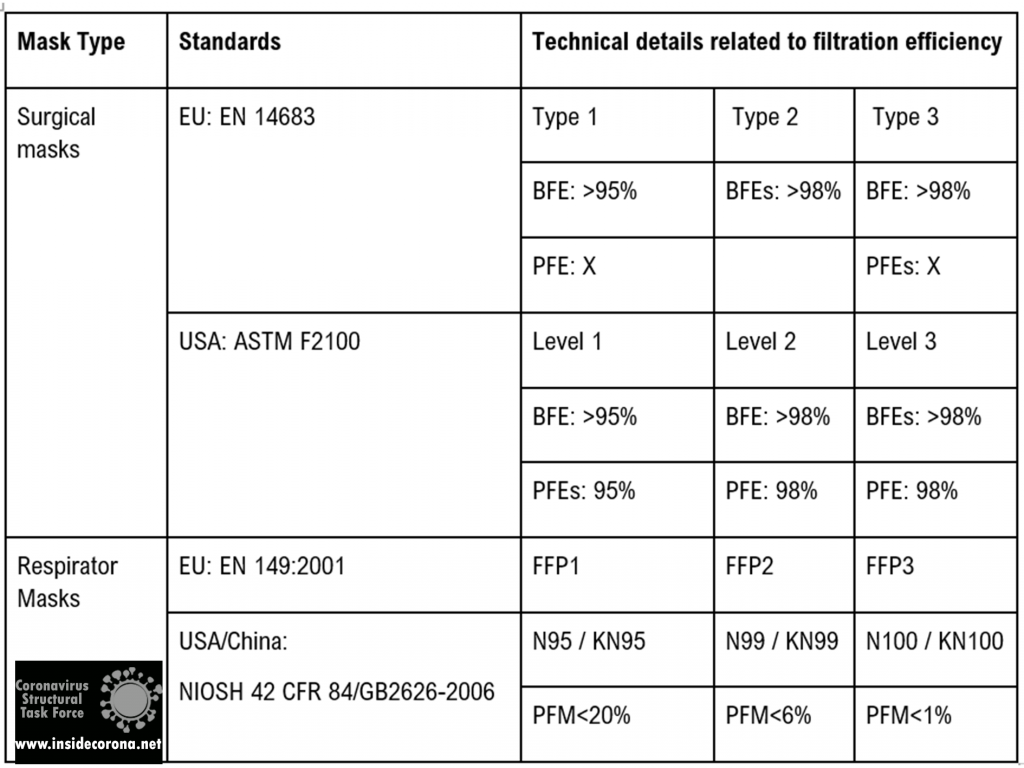
Mechanisms of protection
Masks ensure protection from viral spread in three main ways1,5:
Flow resistance inhibits the momentum of exhaled droplets and the velocity of incoming airborne aerosols. This significantly reduces the risk of infection in the vicinity of an infected person, protecting third parties as well. This is afforded by surgical masks, FFP2/N95/KN95, or better particle filtering respirator masks.
Droplet filtration blocks out large droplets via gravity sedimentation, inertial impaction, and minimizing contact of hand to mouth, nose, or other facial canals with access to the respiratory tract. It is afforded by most kinds of masks.
Aerosol filtration reduces the spread of aerosols via interception, diffusion, and electrostatic attraction. Electrostatic effects likely result in charge transfer with nanoscale aerosol particles. It is afforded by FFP2/N95/KN95 or better particle filtering respirator masks.
At small aerosol droplet sizes in the range of 0.1 to 1 μm, the mask layers prevent particles from passing mainly by blocking movement of particles with the fibers in the filter layer and, hence, not allowing diffusion. For nanometer-sized particles, which can easily slip between the openings in the network of filter fibers, electrostatic attraction is the main way by which mask layers remove low mass particles, which are attracted to and bind to the fibers. This filtering of particles by electrostatic attraction is generally most efficient at low speed of the particles such as the speed of aerosols released by breathing through a face mask.
It is important to note that openings and gaps (such as those between the mask edge and the face) can compromise the performance. Findings indicate that leakages around the mask area can reduce efficiencies by ∼50% or more, pointing out the importance of a proper “fit”8.
Although a home-made fabric mask will at least offer some degree of protection against larger droplets and prevent access to facial features, it will not be very effective in protecting against respirable particles and droplets with a diameter of 0.3 to 2 μm, as these pass through the materials largely unfiltered5.
Thus, the inhalation of droplets containing viruses can be prevented by using a tight-fitting mask with particle filtering properties (self-protection). The FFP2/FFP3 mask type is very well suited to protect people from an infection by means of aerosol even when the environment is strongly contaminated with infectious droplets5.
How does mask structure affect filter particles?
For high filtration and blocking efficiency, the construction of masks layers is very important. Factors that contribute to this efficiency are these4,8:
Movement of droplets/aerosols is directly affected by interfiber spacing of the mask material and the number of layers. Combining layers of differing fiber arrangement to form hybrid masks uses mechanical filtering and may be an effective approach.
Electrostatic interaction impeding aerosol transmission is influenced by the type of mask material. Electrostatic attraction mainly affects the removal of low mass particles, which are attracted to and bind to the fibers. Leveraging electrostatic filtering may be another effective approach8.
The SEM pictures below show the structure and construction of mask fibers and give an insight into the factors that contribute to their high filtering and blocking efficiency.
An FFP2 mask combines layers featuring different spacing and fiber network types to form hybrid masks, employing both mechanical and electrostatic filtering.
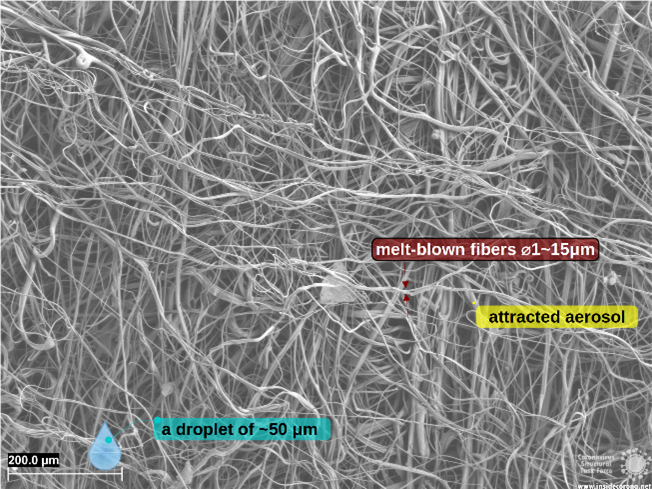

Why are FFP masks superior?
Surgical and respiratory masks are compliant to regulations that guarantee to fulfill certain standards (cf. Table 1). The superior protection of FFP masks stems partially from its filtering layer (cf. Figure 3), using electrostatic filtration to block smaller particles (~0.1 µm).
Conclusion
While maintaining a safe distance from an infected or possibly infected person will prevent spread of infection through direct contact and droplet transmission, maintaining a safe distance may not effectively prevent the spread of infection through airborne aerosols. This is where it becomes very important to wear a mask.
Masks offer self-protection and minimize transmission of potentially infectious exhaled droplets to the surrounding atmosphere. However, in some situations like closed rooms or highly contaminated places, only masks with high blocking and filtration efficiencies will offer this kind of protection, provided they are closely fitted to prevent air from flowing around the mask edges.
The authors would like to explicitly thank Carl Zeiss GmbH, who provided the microscopic images.
References
1. Anand, S. & Mayya, Y. S. Size distribution of virus laden droplets from expiratory ejecta of infected subjects. Sci. Rep. 10, 1–9 (2020).
2. Chirizzi, D. et al. SARS-CoV-2 concentrations and virus-laden aerosol size distributions in outdoor air in north and south of Italy. Environ. Int. 146, 106255 (2021).
3. Lee, B. U. Minimum sizes of respiratory particles carrying SARS-CoV-2 and the possibility of aerosol generation. Int. J. Environ. Res. Public Health 17, 1–8 (2020).
4. Sanchez, A. L., Hubbard, J. A., Dellinger, J. G. & Servantes, B. L. Experimental study of electrostatic aerosol filtration at moderate filter face velocity. Aerosol Sci. Technol. 47, 606–615 (2013).
5. Kähler, C. J. & Hain, R. Fundamental protective mechanisms of face masks against droplet infections. J. Aerosol Sci. 148, (2020).
6. Oct, U. COVID-19 Scienti c Brief : SARS-CoV-2 and Potential Airborne Transmission small particles that can move through the air The term “ airborne transmission ” has a specialized meaning in public health practice respiratory microbes The epidemiology of SARS-Co. 2019–2022 (2021).
7. https://www.bfarm.de/SharedDocs/Risikoinformationen/Medizinprodukte/DE/schutzmasken.html Accessed 21 April 2021.
8. Konda, A. et al. Aerosol Filtration Efficiency of Common Fabrics Used in Respiratory Cloth Masks. ACS Nano 14, 6339–6347 (2020).
COVID-19 vaccines were developed in record time and vaccination exercise is of course ongoing in most countries. Everyone is anxious to see the pandemic come to an end for things to return to normal. As of 23rd April 2021, more than 966 million doses have been administered worldwide [1], however, we have to remember that we do not live in a world where everyone is going to be vaccinated. Two questions are of importance here: Are the current strategies and measures in place adequate to contain the virus fast enough, and are there things that could be done differently for a faster sustainable outcome? To this effect, I state a couple of opinions that I believe are quite essential.
Are there other effective strategies for the COVID-19 vaccination campaign that programme managers can adopt for better vaccination coverage?
We have vaccines that are up to 95% efficacious, but more than that is requires for a well-planned campaign to bring the pandemic to an end. When a campaign is well planned and a good approach is adopted, it helps a lot in yielding an expected outcome. These approaches stated below have not yet been well-practised in the course of this pandemic. I believe it will be beneficial for the vaccination programme managers to adopt them accordingly.
Barrier Analysis
This is a rapid assessment tool that project implementers can use to identify behavioural determinants to know why a promoted behaviour has not been absorbed or adopted. This approach was developed in 1990 by Tom Davis who found it very useful in behaviour change projects [2]. Barrier analysis is necessary considering the context of different countries. Until today, some health organizations have used this approach to have successful and impactful projects [3,4]. This approach involves the use of a structured tool to understand why a particular behaviour has not changed even though a reasonable amount of effort has been put into it. It helps you to know the behavioural determinants of a particular behaviour and what needs to be improved for a better outcome. With this, vaccine uptake can be promoted for better vaccination coverage.
Needs Assessment
Needs Assessment is the collection and analysis of information that relates to the needs of affected populations which will help determine gaps between an agreed standard and the current situation. It helps to determine the key activities for intervention. After the designed questionnaires have been administered, retrieved and analysed, the actual area of focus will be more evident before project implementation. This can help prioritise or improve the services offered to a patient for better acceptability. We have used this approach to implement a health care worker vaccination programme in 2016 and it was very successful and impactful [5]. It helped to determine specific behaviours, activities and actual high-risk areas (apart from the normal official statistics) before implementation.
Health behaviour theories that can also be adopted during implementation
Health Belief Model was developed in the 1950s by a group of U.S. Public Health Service social psychologists who tried to know why some disease detection and prevention programmes had few participants [6,7]. It is based on the construct that people are willing to change if they:
- believe their susceptibility to a particular condition (perceived susceptibility),
- believe that the condition may have serious effects (perceived severity),
- believe the need to take action to reduce any negative effect or its severity (perceived benefits),
- believe they can practice the behaviour without anything stopping them (perceived barriers),
- are exposed to the things that make them remember to do the behaviour (e.g., posters, television advertisements, setting a reminder) (cue to action), or
- believe in their ability to practise the behaviour (self-efficacy)
Theory of Planned behaviour assumes that behavioural intention is the most important determinant of behaviour. It says that behavioural intention is influenced by a person’s attitude towards performing a behaviour and by beliefs about whether individuals who are important to the person approve or disapprove of the behaviour (subjective norm) [7].
I do encourage vaccination project managers to employ any of these models for a fast and better outcome since pieces of evidence have shown the effectiveness on vaccine uptake [8].
How have vaccines been distributed?
As of 23rd April 2021, 01:25 CEST, more than 966 million doses have been administered across 172 countries in the world with the rate of 16.4 million doses daily. This is adequate to vaccinate only 6.3% of the global population [1]. More than 20 countries have not started vaccination at all, and they do not have the vaccines. The vaccine distribution has been so uneven when comparing the high-income and low-income countries. A considerable quantity of vaccines has been administered in the high-income countries and they have also booked more vaccines. As of 19th March 2021, high- and upper-middle-income countries have secured more than 6 million doses out of 8.6 billion expected to be produced. There is a very large margin between these two, high-income countries vaccinate 25 times faster than the low-income countries (ones that have been able to procure few vaccines). In Africa, the majority of the countries have doses that can only be enough for <1% of their total population. For example, Nigeria plans to vaccinate at least 70% of the eligible people aged 18 years and above in four phases within the next two years, but this is impossible with the rate they are going [9]. India (the world’s biggest maker of vaccines) usually supplies vaccines to low-income countries that cannot afford very expensive vaccines [10]. Now, India is experiencing a large COVID-19 surge (since April 2021), and they will have to stop some external supply for their use which will cause more shortage in all these low-income countries. This even makes it more difficult to quell the outbreak globally. It is understandable to prioritise one’s country, but we must remember that eradicating a disease is more than that, it is just “a flight away” and the virus will come back to one’s own country—if not a new variant. The fewer coronavirus cases we have globally, the less likely that new variants will emerge.
How do we maintain protection?
All the approved COVID-19 vaccines will protect you from severe infection, hospitalisation and death irrespective of the efficacy rate. From research, when one is infected with COVID-19 and recovered, the person will gain at least a 6-month protection—it varies from person to person [11–13]. New variants will keep emerging, but I believe that at some point, the variants will not be very strong anymore as vaccination continues. This is why we need to have a running immunization programme at regular intervals. Since we are not yet certain about how long the immunity offered by the vaccine will last and as new variants keep emerging, it will be nice for countries to establish Supplementary Immunization Activities (SIAs), especially for COVID-19.
SIAs is a programme put in place to complement vaccination and get more people vaccinated. It does not replace routine immunization. This is done to boost immunity and prevent emerging variants. In the case of COVID-19, I think it should be carried out annually or on an even shorter schedule.
How do we get more people vaccinated with COVID-19 vaccines?
The best way to reach herd immunity is through vaccination. Vaccination is always considered effective and successful when people are willing to receive the vaccines. You can manufacture the most efficacious vaccine, but when people have disinformation or contrary opinion in getting vaccinated, it is a wasted effort.
Religious and traditional leaders are vital for a successful roll-out of COVID-19 vaccines mostly in the developing world. Generally, these trusted leaders should be involved [4].
Also, mainly in the developed world, one of the major setbacks in vaccination is that people do not want to be told what to do, they see it as a form of violation to their freedom. Adopting the theories mentioned above can help deal with the major determinants of such behaviour.
Incentivising vaccination at this point is important—giving people a quality shirt (for example, with an imprint of their favourite leader) or likes, which they can proudly wear outside, can help spread awareness. When others see that this person has been vaccinated and “did not die”, they can re-consider getting vaccinated.
Other prevention strategies that need to be encouraged other than masking, hand-washing, and social distancing:
In disease prevention, it is always encouraged to observe any safe and effective measures first before thinking of medications or treatments, in order to protect yourself. It is not yet peer-reviewed or strongly proven that the vaccines protect you from spreading or contracting the virus—most especially the new variants—, and that is the reason you are always encouraged to employing masks and social distance. I must encourage these other preventive measures as well.
Nasal irrigation and gargling: Nasal irrigation- is simply the practice of washing your nasal cavity to reduce mucus and germs. There may be a shred of limited clinical evidence on this for SARS-CoV-2 infection, nevertheless, other studies support this, and I agree with them [14,15]. This is usually performed by mixing salt and baking soda in lukewarm water, using it to flush your nostril. Please do keep any device you are using sterile and clean, anyway. A lot of people forget that it is important to wash your nose just as you wash your body mostly to prevent respiratory tract infections. Whenever I take a shower, I wash my nose and expel, it has drastically reduced my chances of getting an allergic reaction.

Figure 1: A person performing nasal irrigation.
The study conducted in 1999 to ascertain the use of isotonic saline nasal irrigation among woodworkers, who face challenges due to the inhalation of dust particles, indicates that it significantly improves nasal symptoms. Also, more than half of the subjects continued to practise it after one year [16].
Gargling with warm saltwater can also help reduce the accumulation of mucus and germs in the upper respiratory tract [17,18].
Eating habit: Poor diet behaviour plays a vital role in disease prevention and management. You may wish to cook more often and stop eating junks or unhealthy food. I believe that eating food enriched with vitamins will help reduce the severity of illness. A well-nourished and hydrated (drink enough water) person is more likely to have a stronger immune response to fight infections.

Figure 2: Unhealthy diets.
Elevator: People must be aware that elevators are not well ventilated. Viral droplets sneezed out can spread fast in there. You may wish to reduce the number of times you use elevators and always wear your masks inside the elevator.
Conclusion
Good strategies and planning are key to a successful vaccination programme, which is very important in tackling the ongoing pandemic. Some of the above-mentioned theories are worth considering for better vaccine uptake. Focusing on your country will not end the pandemic, and it will even affect the low-income countries negatively as they will find it more difficult to recover economically not just from the virus. There may be new variants of SARS-CoV-2 every year or more, but after a while, the variants will be less of a problem to deal with, as people get booster shots yearly for the next two or three years. Only one human virus (smallpox) has been eradicated in history using vaccination [19,20]. Hence, SARS-CoV-2 can be controlled to the point that it will not cause major disruption to our lives, and things will return to normal. Government and policymakers around the world should put in more effort to ensure that vaccines are supplied quickly across the globe.
References
[1] Bloomberg. More Than 966 Million Shots Given: Covid-19 Tracker. Bloomberg.com [Internet]. [cited 2021 Apr 23]; Available from: https://www.bloomberg.com/graphics/covid-vaccine-tracker-global-distribution/
[2] A Practical Guide to Conducting a Barrier Analysis [Internet]. [cited 2021 Apr 20]. Available from: https://pdf.usaid.gov/pdf_docs/PA00JMZW.pdf
[3] World Vision. BA Tabulation Tables and Results Summaries (by country). [Internet]. [cited 2021 May 25]. Available from: https://drive.google.com/drive/folders/1uUb_IWgw6Kk8wgWHFb3Yxr5kpyEDJaMk?usp=sharing
[4] Harris N. Faith leaders must play key role in COVID-19 vaccine roll-out [Internet]. World Vision. [cited 2021 Apr 20]. Available from: https://www.worldvision.org/about-us/media-center/faith-leaders-must-play-key-role-in-covid-19-vaccine-roll-out
[5] Arogundade L, Akinwumi T, Molemodile S, Nwaononiwu E, Ezika J, Yau I, et al. Lessons from a training needs assessment to strengthen the capacity of routine immunization service providers in Nigeria. BMC Health Serv Res. 2019 Sep 14;19(1):664.
[6] Wong MCS, Wong ELY, Huang J, Cheung AWL, Law K, Chong MKC, et al. Acceptance of the COVID-19 vaccine based on the health belief model: A population-based survey in Hong Kong. Vaccine. 2021 Feb 12;39(7):1148–56.
[7] National Cancer Institute- Theory at a Glance [Internet]. [cited 2021 Apr 26]. Available from: https://cancercontrol.cancer.gov/sites/default/files/2020-06/theory.pdf
[8] Chu H, Liu S. Integrating health behavior theories to predict American’s intention to receive a COVID-19 vaccine. Patient Educ Couns [Internet]. 2021 Feb 17 [cited 2021 May 4]; Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7889032/
[9] Joint press statement by NPHCDA, WHO and UNICEF on the arrival of COVID-19 vaccine in Nigeria [Internet]. [cited 2021 May 26]. Available from: https://www.unicef.org/nigeria/press-releases/joint-press-statement-nphcda-who-and-unicef-arrival-covid-19-vaccine-nigeria
[10] Serum Institute of India. About Serum Institute Of India Pvt. Ltd. [Internet]. [cited 2021 May 8]. Available from: https://www.seruminstitute.com/about_us.php
[11] Hall V, Foulkes S, Charlett A, Atti A, Monk EJM, Simmons R, et al. Do antibody positive healthcare workers have lower SARS-CoV-2 infection rates than antibody negative healthcare workers? Large multi-centre prospective cohort study (the SIREN study), England: June to November 2020. medRxiv. 2021 Jan 15;2021.01.13.21249642.
[12] Dan JM, Mateus J, Kato Y, Hastie KM, Yu ED, Faliti CE, et al. Immunological memory to SARS-CoV-2 assessed for up to 8 months after infection. Science [Internet]. 2021 Feb 5 [cited 2021 Apr 26];371(6529). Available from: https://science.sciencemag.org/content/371/6529/eabf4063
[13] Abu Raddad LJ, Chemaitelly H, Malek JA, Ahmed AA, Mohamoud YA, Younuskunju S, et al. Assessment of the risk of SARS-CoV-2 reinfection in an intense re-exposure setting [Internet]. Epidemiology; 2020 Aug [cited 2021 May 26]. Available from: http://medrxiv.org/lookup/doi/10.1101/2020.08.24.20179457
[14] King D, Mitchell B, Williams CP, Spurling GK. Saline nasal irrigation for acute upper respiratory tract infections. Cochrane Database of Systematic Reviews [Internet]. 2015 [cited 2021 Apr 26];(4). Available from: https://www.cochranelibrary.com/cdsr/doi/10.1002/14651858.CD006821.pub3/full
[15] Panta P, Chatti K, Andhavarapu A. Do saline water gargling and nasal irrigation confer protection against COVID-19? Explore (NY). 2021;17(2):127–9.
[16] Rabone SJ, Saraswati SB. Acceptance and effects of nasal lavage in volunteer woodworkers. Occup Med (Lond). 1999 Aug;49(6):365–9.
[17] Ramalingam S, Graham C, Dove J, Morrice L, Sheikh A. Hypertonic saline nasal irrigation and gargling should be considered as a treatment option for COVID-19. J Glob Health [Internet]. [cited 2021 Apr 26];10(1). Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7193539/
[18] Satomura K, Kitamura T, Kawamura T, Shimbo T, Watanabe M, Kamei M, et al. Prevention of upper respiratory tract infections by gargling: a randomized trial. Am J Prev Med. 2005 Nov;29(4):302–7.
[19] Greenwood B. The contribution of vaccination to global health: past, present and future. Philos Trans R Soc Lond B Biol Sci [Internet]. 2014 Jun 19 [cited 2021 Apr 26];369(1645). Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4024226/
[20] World Health Organization. Smallpox [Internet]. [cited 2021 Apr 26]. Available from: https://www.who.int/health-topics/smallpox#tab=tab_1[
There is a secret code that virologists use to talk about the new coronavirus. This code is made up of synonymous words and abbreviations for each of the 28 proteins which facilitate the viral life cycle. In this article, we will shed some light on this mythical language.
First of all, SARS-CoV-2 has three classes of proteins:
Structural Proteins, namely the spike protein, the membrane protein and the envelope protein as well as the nucleocapsid, which forms an extra shell around the single-stranded RNA, are also known as the S-, M-, E- and N-Protein.
Non-structural proteins (NSP) ensure the viral life cycle but are not making up the hull or nucleocapsid; These are conveniently numbered 1–16.
And then there are accessory proteins, which seem to be more important in-vivo than in-vitro, and most of them have not yet been structurally determined.
Of course, this nice and clear naming scheme tells you little about the function and properties of the different proteins, which is why virologists invented plenty of other names for them. And this is where the confusion begins.

NSP3, for example, contains two ubiquitin-like (UBL1 and UBL2) domains, a papain-like protease (PLpro, PL2pro) domain (which includes a zinc finger), a "macro" domain (also known as X domain, Mac1, or ADP ribose phosphatase), a hypervariable region (also called Glu-rich acidic domain or HVR), two transmembrane domains (TM1 and TM2), an ecto (3Ecto) domain (which is also a zinc finger), a conserved domain of unknown function called Y1, and a coronavirus-specific carboxyl-terminal (CoV-Y) domain. The SARS-unique domains, or SUDs—namely SUD-M, SUD-N, and SUD-C—were all renamed after it was found out they are not unique to SARS: SUD-N is now Mac2, SUD-M is Mac3 and SUD-C is called DPUP.
If this was not enough to convince you that all of this is confusing, here are some additional names:
Spike: S-Protein, surface glycoprotein, E2 glycoprotein
NSP1: leader protein
NSP5: 3CLpro, SARS-CoV-2 3C-like protease, 3C-like proteinase, main protease, NSP5A_3CLpro, NSP5B_3CLpro, Mpro, Non-structural protein 5
NSP9: Non-structural protein 9, ssRNA-binding protein
NSP10: Non-structural protein 10, growth factor-like protein, GFL
NSP12: RNA Polymerase, RNA-dependent RNA Polymerase, NiRAN, RdRp
NSP13: NSP13-pp1ab, non-structural protein 13, helicase, NTpase, Hel
NSP14: NSP14A2_ExoN, SARS-CoV-2 3'-to-5' exonuclease, non-structural protein 14, NSP14B_NMT
NSP15: NSP15-A1, SARS-CoV-2 endoRNAse, NSP15B-NendoU, NendoU, uridylate-specific endoribonuclease NendoU
All these names are certainly hard to remember, but as a scientist you need them in order to save the world! So, we made a handy glossary for you that you can access here.
If you have any more suggestions or corrections for the glossary, please let us know in the comments!
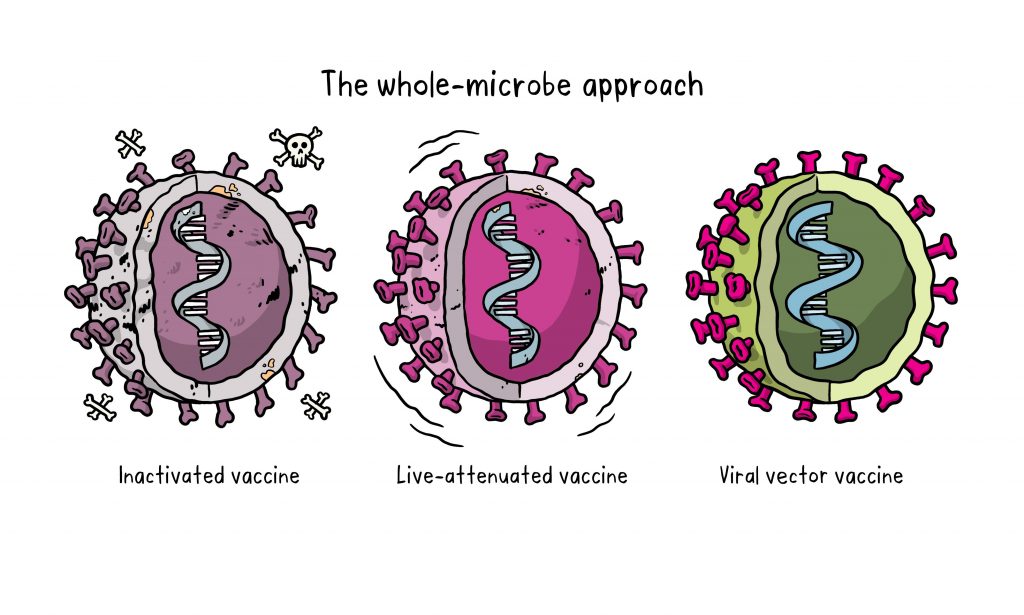
In addition to mRNA vaccines, another type of vaccine is employed against COVID-19: Vector vaccines—like the one from AstraZeneca—contain a mostly functioning virus. But how do they work exactly? What are their strengths and weaknesses? And finally, are they safe?
The novel mRNA vaccines against SARS-CoV-2 have caused some controversy, and many concerns have been raised. According to these, the vaccine has not been sufficiently tested, long-term effects are yet unknown, the vaccine might affect the patient’s genetic material.
While mRNA vaccines enfold the genetic material in a tiny lipid particle, so-called viral vector vaccines put it in a viral shell that functions as a transporter (vector). A large number of vaccines of this type are under development, and they are all based on various strains of the same virus known for causing the common cold. The European Medicines Agency (EMA) recommended the Oxford–AstraZeneca vaccine for approval in late January, and the United States approved the use of a similar vaccine from Johnson & Johnson in February. Why?
Viruses are transporters for genes
Viruses are not considered living organisms because they have no metabolism of their own. They contain only the genes that cause the host cell to produce new viruses.
In the evolution of viruses, some of their skills have become exceptionally refined. Viruses can specifically infect host cells, escape the host’s immune defenses, and insert their genetic material into the host cell. Once molecular biologists recognized these capabilities, scientists could turn viruses into tools that do not cause disease through genetic engineering. As modified tools, they are referred to as viral vectors.
Viruses as tools
To make viruses useful for research purposes, they are modified. Their natural genes must be changed or deleted. What is left in the end is a transporter that can be used to introduce desired genes into a host. This method is used, for example, to produce transgenic (genetically modified) plants or cell lines. In this process, researchers usually keep some natural functions of the virus such as the capability to invade host cells.
Viruses as vaccines
What exactly must be changed to turn a virus into a vaccine?
In order not to cause a disease, the virus must lose its ability to multiply inside the body. Viruses without the ability to reproduce are called "replication-deficient". In the past, replication was disabled via lengthy cell culture techniques but, since the 2000s, genetic engineering has made it possible to selectively remove or modify genes of a virus to stop it from reproducing. Removing critical genes can completely prevent reproduction of the virus and any chance of reversion to the pathogenic (disease-causing) variant can be prevented [3].
Gene modification can also be used to introduce structures such as the spike gene of SARS-CoV-2 into a virus that is harmless to humans. This addition turns the virus into a vaccine.
Viruses have been used as tools for a long time. Viral vectors were first described in 1972 [1]. In the early 1990s, gene transfer was first used for therapy through an attenuated adenovirus as vector [2].
Across the last twenty years, a number of vaccines based on viruses have been developed. An example of an approved vector vaccine is the Ebola vaccine Ervebo [4]. Because there is no treatment for Ebola to date, the epidemics from 2013 to 2020 were devastating for Central Africa. Mortality rates ranged from 25 to 90% [5]. In August 2018, Ervebo was used for the first time to vaccinate a large group of people. It proved to be highly effective [6] and was also licensed for employment in the EU in 2019 [5].
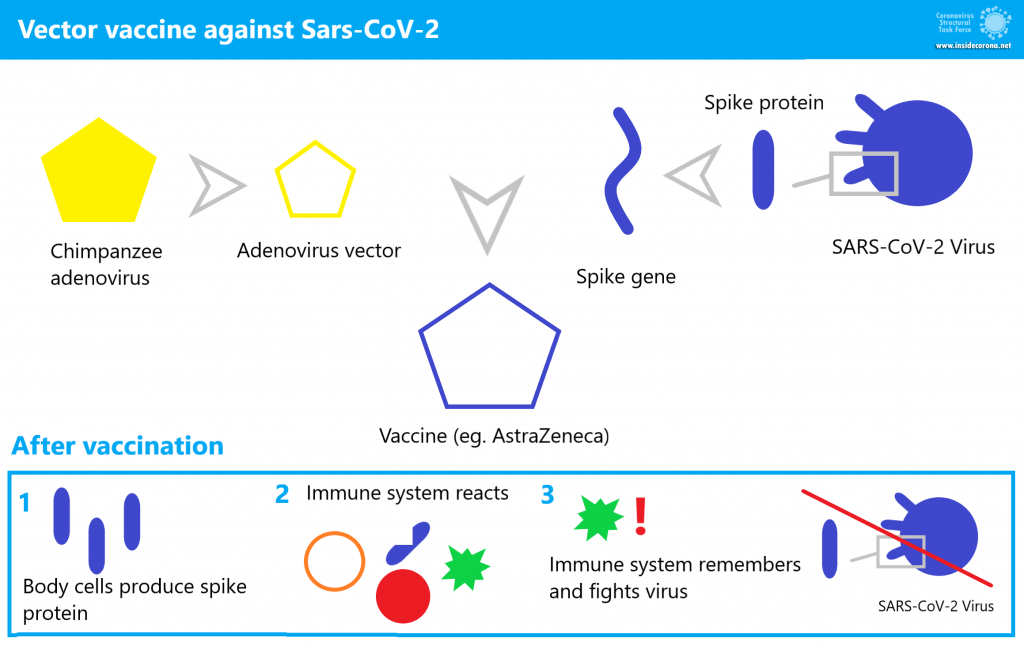
How does the AstraZeneca vaccine work?
This vaccine is known as AZD1222 (also called ChAdOx1 nCoV-19) [8]. It is an adenovirus that is similar to a pathogen for the common cold, but this strain was originally found in chimpanzees. It has been modified to be replication-deficient. AZD1222 was also modified to contain the coronavirus’ surface protein (spike) gene [9]. This chimpanzee virus is used because most people’s immune systems are familiar with adenoviruses that infect humans, and their immune systems might respond to those viruses before they can infect cells and produce spike protein. In comparison, the chimpanzee virus is unfamilar to the human immune system and is hence capable of infecting a cell [10]. Side effects can occur, but they do not stem from the actual disease. Feeling unwell after being vaccinated is triggered by our immune system fighting against an intruder.
AZD1222 invades some human host cells just like a usual common cold virus. During this process, the vector transports the gene of the spike protein into the host cell. The spike DNA is utilized by the cell to produce spike protein. This spike protein acts as an antigen, a substance that triggers an immune response. The immune system recognizes it as foreign and starts attacking, antibodies are produced and the few "infected" cells containing AZD1222 genetic material are destroyed. Antibodies bind to the antigen like pieces of a puzzle and hold on to it. The antigen-antibody clumps are then degraded.
What remains from the whole process are memory cells that recognize the spike protein when attacked by the real SARS-CoV-2. Previous training with the vaccine makes the next immune response faster and more effective. This prevents disease before it can break out.
How effective and safe is the vaccine?
AZD1222 has already been tested in over 20,000 people across three phases of clinical trial [11]. Since its approval, data has been continued to be collected and combined into additional studies. Through these, it is possible to investigate possible side effects even more thoroughly since millions of people are being vaccinated and observed in the "real world".
One example is a British study on efficacy in people over 70 years of age. The result: A single dose of AstreZeneca’s viral vector vaccine or BioNTech’s mRNA-based vaccine is effective against a symptomatic Corona infection in 60-75% of cases. The likelihood of hospitalization is reduced by 80% with either vaccine [12].
The first nationwide study took place in Scotland. It was led by the University of Edinburgh and collected data from 5.4 million vaccinated people. Preliminary data show that AstraZeneca’s single-dose vaccine prevented a severe course—requiring hospitalization—in up to 94% of cases [13].
Through these real world studies, it is now known that the vector vaccine shows greater effectiveness with a larger interval between the first and second dose. Therefore, these studies recommend an interval of twelve weeks [14].
In terms of side effects, some data has already been collected on AZD1222 in the UK due to the earlier start of vaccination [15]. Common side effects experienced by one in ten patients receiving vaccination include:
- tenderness, pain, warmth, itching, or bruising at the injection site,
- feeling generally unwell or tired,
- chills or feverish feeling,
- headache,
- nausea, and
- joint pain or muscle soreness.
These frequent side effects are similar to those of the BioNtech vaccine. However, no serious side effects have been detected in the trials. Fever or flu-like symptoms are less common and are a general sign of the vaccine activating the immune system and therefore taking effect. In any case, these symptoms are not comparable to the risks of a severe COVID-19 course.
The EMA is currently looking into cases of blood clotting that occurred shortly after an AstraZeneca vaccination. So far there are no indications that the vaccine is unsafe. The frequency of blood clots in vaccinated people is not higher than it generally is in the population. According to AstraZeneca [15.1] the risk of pulmonary embolism, deep vein thrombosis (DVT) or thrombocytopenia is not higher in people receiving the vaccine compared to the general public. Among 17 million vaccinated patients in Europe, some are expected to suffer from these diseases, but it is very likely not linked to the vaccine.
What is in the vaccine?
In addition to the modified adenovirus, the vaccine contains several other substances [16].
Most are additives that stabilize the vaccine and make administration easier. These include amino acids, stabilizers, alcohol, sugar, salt, binders, and water. In addition, the vaccine is free of food allergens (such as soy or lactose) and contains no ingredients of human or animal origin. That sounds incredible for a chimpanzee virus that needs a host to reproduce. How can this be?
All modifications and amplification of the virus took place in a human cell line used in the laboratory for such purposes, so-called HEK293 cells. These cells are, however, not part of the vaccine.
What are the advantages of this type of vaccine?
Viral vector vaccines—just plain RNA/DNA vaccines—can be developed very quickly. The reason: As soon as the genes of the pathogen are known, they can be used for vaccine development. This shortens the time until clinical trials can begin. This makes vector vaccines suitable for sudden epidemic outbreaks [18].
One advantage of the AstraZeneca vaccine is its dosage. Oxford–AstraZeneca initially tested different dosing approaches for their vector vaccine. Vaccinations with two standard doses at intervals of four to twelve weeks and a vaccination with only one dose were compared. For this, a standard dose contains 5x1010 virus particles [19].
"By using a more effective dosing regimen," said Professor Pollard, the lead scientist at Oxford, "more people could be served with the same amount of vaccine."[20]. The ideal regimen was found to be the administration of a half dose (2.2x1010 virus particles) followed by a standard dose at least one month apart. The efficiency in this case was 90% [21]. However, this must first be confirmed by additional data. So far, vaccination is done with two standard doses.
Conclusion
In nature, viruses have perfected the ability to insert their genetic material into hosts. The fascinating abilities of viruses can nowadays be used by scientists. They are therefore used to create genetically modified plants or cells, to treat hereditary diseases, and to produce vaccines.
Vector vaccines—such as AstraZeneca’s—contain harmless and non-reproducing viruses that contain a part of a disease-causing virus. By artificially "infecting" the patient with the vector vaccine, the immune system is trained to respond to the pathogen and can react more quickly and effectively in the event of a real infection in the future.
[1] Jackson D.A., Symons R.H., Berg P. Biochemical method for inserting new genetic information into DNA of Simian Virus 40: Circular SV40 DNA molecules containing lambda phage genes and the galactose operon of Escherichia coli. Proc. Natl. Acad. Sci. USA. 1972;69:2904–2909. doi: 10.1073/pnas.69.10.2904.
[2] Zabner, Joseph et al. 1993 Adenovirus-mediated gene transfer transiently corrects the chloride transport defect in nasal epithelia of patients with cystic fibrosis Cell, Volume 75, Issue 2, 207 – 216.
[3] Using directed attenuation to enhance vaccine immunity, Rustom Antia, Hasan Ahmed, James J Bull, bioRxiv 2020.03.22.002188; doi: https://doi.org/10.1101/2020.03.22.002188
[4] https://www.who.int/vaccine_safety/committee/topics/ebola/Jul_2019/en/
[5] https://www.ema.europa.eu/en/news/first-vaccine-protect-against-ebola
[6] https://www.thelancet.com/journals/lancet/article/PIIS0140-6736(16)32621-6/fulltext#seccestitle160
[7] https://www.nature.com/articles/d42859-020-00016-5
[8] https://www.ema.europa.eu/en/documents/product-information/covid-19-vaccine-astrazeneca-product-information-approved-chmp-29-january-2021-pending-endorsement_en.pdf
[9] https://jvi.asm.org/content/jvi/77/16/8801.full.pdf
[10] https://www.cdc.gov/vaccines/covid-19/hcp/viral-vector-vaccine-basics.html
[11] https://clinicaltrials.gov/ct2/show/NCT04516746
[12] https://www.medrxiv.org/content/10.1101/2021.03.01.21252652v1
[13] https://www.ed.ac.uk/files/atoms/files/scotland_firstvaccinedata_preprint.pdf
[14] https://www.rki.de/DE/Content/Kommissionen/STIKO/Empfehlungen/AstraZeneca-Impfstoff.html
[15] https://www.gov.uk/government/publications/regulatory-approval-of-pfizer-biontech-vaccine-for-covid-19/information-for-uk-recipients-on-pfizerbiontech-covid-19-vaccine#side-effects
[15.1] https://www.astrazeneca.com/media-centre/press-releases/2021/update-on-the-safety-of-covid-19-vaccine-astrazeneca.html
[16] https://www.astrazeneca.at/content/dam/az-at/pdf/2021/Vaccine%20guide%20for%20HCPs%20-%202021-02-02.pdf
[17] Draper, S., Heeney, J. Viruses as vaccine vectors for infectious diseases and cancer. Nat Rev Microbiol 8, 62–73 (2010). https://doi.org/10.1038/nrmicro2240
[18] https://www.frontiersin.org/articles/10.3389/fimmu.2018.01963/full
[19] https://papers.ssrn.com/sol3/papers.cfm?abstract_id=3777268
[20] https://www.astrazeneca.com/media-centre/press-releases/2020/azd1222hlr.html
[21] https://www.thelancet.com/journals/lancet/article/PIIS0140-6736(20)32661-1/fulltext
Introduction:
Vaccination is a great means of achieving public protection against diseases. The main goal of any vaccine manufacturer is to produce a vaccine that will be safe and effective in preventing the target disease. Before any vaccine is rolled out for mass vaccination or campaign, it must have met the required rigorous scientific and ethical standards, ensuring safety, efficacy, purity, and potency (1). A vaccine is considered safe and effective when used correctly, leading to a successful vaccination campaign. However, from the general health point-of-view, vaccines are not risk-free and there are occasionally Adverse Events Following Immunization (AEFI) (2). The risk is of course outweighed by the benefits of vaccines as they protect us from vaccine-preventable diseases. Here, I will discuss the safety measures vaccination undergoes for it to be a success.
What does vaccination mean?
Vaccination is a simple, safe, and effective way of protecting people against harmful diseases, before they come into contact with them (3).
What is vaccination safety?
This entails the absence of preventable harm to the recipient, health care worker and the society at large during and after vaccination.
What are the stages of vaccine development before vaccination?
Vaccine development starts with the assessment of public health needs and priorities which progresses to scientific research on the target disease, pre-clinical and eventually, clinical trials (4). The following are the stages of vaccine development:
Research stage; involves the identification and isolation of the target antigen.
Pre-clinical stage; the potential vaccine is being tested on cells and animals after a careful examination.
Clinical trials (three phases); if the vaccine is safe in animals, it will then be suggested to be tested in human volunteers.
The clinical trials are usually in three phases.
- Phase 1; trials involve a smaller number of people of about twenty to hundred healthy volunteers, this is designed to test and determine vaccine safety, doses, immune response (immunogenicity) and route of administration.
- Phase 2; starts after the successful completion of phase 1. This may involve hundreds of volunteers with double-blind (the researchers and the research volunteers do not know if the research volunteers are in the vaccine group or the placebo group, to prevent bias) studies with a placebo-control group. It further tests safety and the amount of dose to provoke an immune response.
- Phase 3; hundreds to thousands of volunteers are given the vaccine in comparison with the placebo group. It involves a randomized double or single-blind (the research volunteers do not know whether they are in the vaccine group or the placebo group, to prevent bias), placebo-controlled group (inactive substance, sometimes another type of vaccine is given to the research volunteers) to evaluate more on the vaccine safety, efficacy and side effects.
This is then followed by regulatory review, approval and manufacturing. After the product is rolled out, a surveillance system is established for continuous monitoring of the vaccine, immunization coverage and possible adverse reactions for risk-management (4,5). COVID-19 vaccines underwent these processes as well.
Who is responsible for vaccine approval?
Following scientific, ethical and international standards, approval on clinical trials, results, and licensing are done by the national and regional regulatory authorities in the countries where vaccines are manufactured (6). This must be guided by principles of fairness, transparency and accountability to ensure safety.
How are vaccines stored and handled?
Cold chain, sometimes referred to as “Supply chain” is the system used for storing vaccines in good condition. In the cold chain system, the levels designed to keep vaccines within recommended temperature ranges, from the point of manufacture to the point of administration differ. This means that the storage temperature range for every level may differ provided the potency of the vaccine is maintained. Vaccines can be sensitive to freezing or light but all vaccines are sensitive to heat. This is what the Vaccine Vial Monitor (VVM) on the vial helps to monitor. VVM is a chemical indicator label attached to the vaccine container (vial and ampoule) by the vaccine manufacturer which helps to monitor the temperature of a vaccine and ensure that no heat-damaged vaccine is administered. If the colour of the inner square is the same colour or darker than the outer circle, the vaccine has been exposed to too much heat and should be discarded. All vaccines must be discarded after the expiry date or six hours of opening. The recommended diluent is used for the reconstitution of freeze-dried vaccines before use.
Are some vaccines made for some persons?
Vaccination is carried out based on dosage, age group or sex. This is because there is a specific group of people used during the clinical trials who will determine the first target group to be vaccinated when the vaccine is rolled out. Notwithstanding, other group of people may still receive the vaccine as the vaccination goes on once safety is ensured. Please always check the fact sheet for any vaccine you wish to receive on the manufacturer`s website for such updates.
How are injection devices disposed of after a vaccination session?
Injection equipment can pose danger to the environment if not well disposed of, leading to environmental pollution. When needles and syringes are thrown into the bodies of water, they cause contamination to the environment and injury to the wildlife. Safety boxes are sharps waste containers that needles cannot penetrate, they are used to keep the used needles and syringes temporarily during a vaccination session. It is disposed of immediately after vaccination sessions to maintain safety. Appropriate use of a safety box is necessary to avoid needle-stick injuries (when the skin is pricked by needles accidentally) to the health care worker or other individuals. Special incinerators are used to burn these needles and syringes to minimise the toxic release into the environment.
Source: Adobe Stock

Source: Adobe Stock
What is Adverse Events Following Immunization (AEFI)?
This is any untoward medical occurrence which follows immunization and which does not necessarily have a causal relationship with the usage of the vaccine. AEFI can be vaccine product-related, vaccine quality defect-related, vaccination error- related, vaccination anxiety-related or a coincidental event.
Some minor vaccine reactions-AEFIs, you need to know:
- Pain, redness, swelling at the site of injection
- Irritability, malaise
- Slight fever
- Slight headache
- Mild muscle pain
- Loss of appetite
- Joint pain
- Lymphadenopathy (swollen lymph nodes, it can be under the armpit in the same arm of injection, etc.)
Some rare and severe vaccine reactions-AEFIs, you need to know:
- Febrile seizures (convulsions that occur usually in children as a result of high body temperature)
- Thrombocytopenia (low platelet count in the blood which usually helps to stop bleeding when one gets injured)
- Anaphylaxis (severe allergic reaction)
- Sterile abscess (swelling in the injection site usually occurs when an injection is not completely absorbed into the skin which makes pus build up in the tissue)
- Difficulty in breathing
- Encephalopathy (damage to the brain, affecting one`s mental state)
- Persistent inconsolable crying or screaming
Severe AEFIs are rare, however, every AEFI must be reported to the health care worker or appropriate authority.
Before receiving any vaccine, please discuss with the vaccination provider about all of your medical conditions such as:
- past and present allergic reactions
- current fever
- if you are taking blood thinner medications
- if you have any bleeding disorder
- if you are immunocompromised
- if you are on a medication that affects your immune system
- if you are pregnant or plan to become pregnant very soon
- if you are breastfeeding
- your previous vaccination history (7,8).
What does surveillance (pharmacovigilance) mean?
Pharmacovigilance is a part of the surveillance system designed to detect, assess, understand, respond and preventing adverse drug reactions, including reactions to vaccines-AEFIs in every country. Both national and international levels have surveillance systems for good monitoring and immediate actions in response to AEFIs. This helps to ensure the safety of vaccines even as vaccination is ongoing (2).
Conclusion:
Vaccines follow many rigorous scientific processes before and after being approved to ensure safety and effectiveness. Vaccination is safe when no harm is posed to the patient, health worker and society (avoiding pollution and injuries through proper disposal of injection wastes). If a vaccine requires a subsequent dose, you must receive the same type of vaccine as the initial dose. Always keep your vaccination card for the next visit and follow your vaccination schedule. It is encouraged to not go for a second dose of vaccine if you had a serious allergic reaction or side effect after the first dose. Discuss your medical history with the vaccination provider before taking a vaccine. Please wait for some time after receiving your vaccine for a little safety monitoring before going home. Vaccines work!
References:
1. Centers for Disease Control and Prevention. U.S. Vaccine Safety - Overview, History, and How It Works | CDC [Internet]. 2020 [cited 2021 Mar 4]. Available from: https://www.cdc.gov/vaccinesafety/ensuringsafety/history/index.html
2. World Health Organization. WHO Vaccine Safety Basics [Internet]. [cited 2021 Mar 3]. Available from: https://vaccine-safety-training.org/overview-and-outcomes-1.html
3. Vaccination_World Health Organization. Vaccines and immunization: What is vaccination? [Internet]. [cited 2021 Mar 3]. Available from: https://www.who.int/news-room/q-a-detail/vaccines-and-immunization-what-is-vaccination
4. Centers for Disease Control and Prevention. Ensuring Vaccine Safety | CDC [Internet]. 2020 [cited 2021 Mar 3]. Available from: https://www.cdc.gov/vaccinesafety/ensuringsafety/index.html
5. Mitchell VS, Philipose NM, Sanford JP. Stages of Vaccine Development_Institute of Medicine (US) Committee on the Children’s Vaccine Initiative: Planning Alternative [Internet]. The Children’s Vaccine Initiative: Achieving the Vision. National Academies Press (US); 1993 [cited 2021 Mar 3]. Available from: https://www.ncbi.nlm.nih.gov/books/NBK236428/
6. World Health Organization_ Vaccine Safety. Vaccines and immunization: Vaccine safety [Internet]. [cited 2021 Mar 3]. Available from: https://www.who.int/news-room/q-a-detail/vaccines-and-immunization-vaccine-safety
7. Moderna. Emergency Use Authorization (EUA) | Moderna COVID-19 Vaccine [Internet]. https://www.modernatx.com/. [cited 2021 Mar 16]. Available from: https://www.modernatx.com/covid19vaccine-eua/
8. Pfizer-BioNTech. Pfizer-BioNTech COVID-19 Vaccine | cvdvaccine.com [Internet]. [cited 2021 Mar 16]. Available from: https://www.cvdvaccine.com
