Introduction
The coronavirus pandemic hit the entire world and caused millions of deaths. More than fifty companies race towards developing a vaccine to stop the disease (1). Vaccination presents a lasting solution to this unfavourable situation, reducing the burden of Coronavirus (2). The first vaccine to be approved for emergency use is an mRNA-based vaccine (3,4). How does it work? Here, I will shed some light on this, but before we go into details, we need to discuss a little about vaccines and how a foreign substance provokes an immune response in the body (immunogenicity).
Early impact of vaccines on humanity
Vaccines have existed since 1796 and have saved millions of lives over the years (5,6). Vaccines need to be given orally, intramuscularly or subcutaneously to, stimulate the body for an immune response and generate lasting immunity.
How the body fights diseases
The human body works very hard to remove any foreign substance (potential pathogens). When pathogens, such as bacteria or viruses enter the body, they attack and spread, causing disease. The immune system fights these infections recognizing a part of the pathogen using white blood cells which consist primarily of macrophages, B-lymphocytes and T-lymphocytes (7). Macrophages are antigen-presenting cells that engulf pathogens and digest them. Macrophages present parts of the pathogens to the T-lymphocytes and B-lymphocytes. When the B-lymphocytes are activated, they produce antibodies that target specific pathogens and destroy them. Also, when the T-lymphocytes are activated by the antigen-presenting cells, they seek out and destroy any infected cell in the body. However, it takes a couple of days for the body to generate the antibodies to fight an infection. After recovering from an infection, the immune system can remember how it fought the infection and can quickly in the future, prevent reinfection by the same or similar diseases. When the infection is gone, the body is left with the supply of memory cells called T-lymphocytes and B-lymphocytes that are responsible for this rapid intervention (7). Vaccines cause the same process but without a serious illness and often even without an infection. Individuals react to vaccines differently, some persons experience symptoms like swelling or redness at the injection site, low-grade fever, tiredness etc. These minor symptoms result as the body tries to build immunity against the disease, and it is your body`s healthy response. It would be no good and impracticable for the vaccine to cause the full-blown disease - instead, vaccines prime the immune system to jump into action should you ever encounter the real thing.
Traditional vaccines
Traditional vaccines contain either an attenuated (live or weakened) virus, an inactivated virus, or even a piece of a viral protein that can produce the immune answer – a so-called antigen that does not cause an actual disease but still stimulates the body to produce antibodies to fight a real infection (8). It tricks the immune system into thinking that an infection has occurred and the immune system responds by producing antibodies against the virus (9). mRNA-vaccines, however, do not contain dead or live pieces of the virus – they contain the genetic instructions necessary for our cells to make copies of the antigen itself. So, what is mRNA?
What is mRNA?
mRNA is how our body encodes information from the genome that is to be used as blueprints to make proteins: In every human cell, we find a nucleus that contains our genome in the form of DNA. This DNA is transcribed into mRNA, which then can leave the nucleus and enter the rest of the cell. DNA itself cannot leave the nucleus and this protects the cell’s genome from damage and manipulation. Once outside the nucleus, the mRNA is translated into proteins which then fulfil all kind of work tasks in our cells – they make up our hair, break down our food, transport oxygen around our body (10,11). They pretty much do everything that makes us go.
An mRNA vaccine exploits this process: When the mRNA vaccine enters our cells, it effectively skips the transcription step (DNA to RNA) and goes straight to translation (mRNA to proteins) to produce antigenic proteins (10,12).
How do the SARS-CoV-2 mRNA-based vaccines work?
The best antigen in the coronavirus SARS-CoV-2 – the bit which is recognized by the immune system – is the spike protein. The outside of the SARS-CoV-2 (Coronavirus) is carries these spike proteins, which the virus uses to enter human cells and cause an infection.
New mRNA vaccines contain messenger RNA which encodes the spike protein and as it enters the cell, the spike proteins are produced within the cell. The spike protein then migrate to the surface of the cell, where the immune system recognizes it as foreign and the body will produce antibodies to fight the infection. At this point, the process to produce immunity is the same as for any other vaccine. When infected again, the body can rapidly supply memory cells- T-lymphocytes and B-lymphocytes that remember how to attack a similar pathogen. So, when the real coronavirus SARS-CoV-2 (spike proteins and all) enters the body, the immune system remembers how it fought the formerly produced spike proteins and can easily fight off the infection.
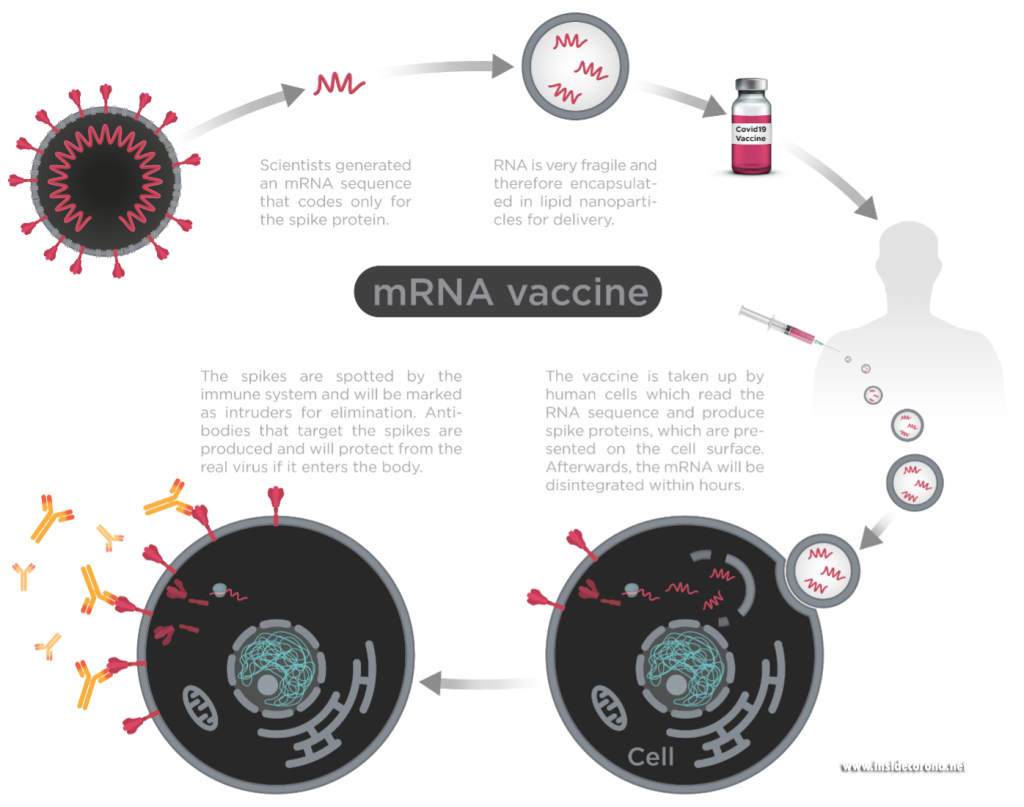
Figure1: Process of mRNA vaccines in the body.
Image by Thomas Splettstösser.
What is in the vaccine and why do we use it?
mRNA-based SARS-CoV-2 vaccines contain the mRNA strands encapsulated in lipid nanoparticles (think soap bubble) to protect it from hot temperatures as well as degrading enzymes (13,14). After being injected intramuscularly, the protection offered by the lipid nanoparticles helps the mRNA to remain active/potent until it enters the cells and releases mRNA.
There are some advantages to mRNA vaccines, and I think it can be expected that we will soon see more of them for other illnesses. The production of mRNA vaccine doses is much faster and cheaper than traditional vaccines since it does not require a long process of growing viral proteins in a cell or an egg which then needs to be deactivated or killed to produce the vaccine (15). Also, in mRNA vaccines there is never a complete virus, so contracting the disease is impossible, which can happen with live vaccines. Finally, should mutations occur, it is relatively straightforward to change the mRNA vaccine to contain this new mutation.
However, the storage temperature of an mRNA vaccine is a challenge: they need to be stored at a very low temperature to maintain its potency until it is ready to be used and this type of freezer is not available everywhere and makes transport difficult compared to a traditional vaccine (14,16).

Figure2: Sample storage freezer for mRNA vaccines.
Image by Yunyun Gao.
Can an mRNA-based vaccine change my DNA?
As the vaccine contains no so-called “reverse-transcriptase”, the spike mRNA cannot be converted into DNA. mRNA-based vaccines cannot even enter the cell’s nucleus. Hence, they are not able to change your DNA. They are just a messenger to produce the spike protein and the enzymes of the cell readily destroy them shortly afterwards. Also, the unfortunate cell itself is destroyed by the immune system once it displays the spike on the surface and the immune system has learned to recognize it.
Do I still need to wear masks?
Also, remember that after getting vaccinated, your body needs time to develop antibodies in sufficient quantities, so you can still contract Coronavirus even after being vaccinated. Hence, we need to maintain safety measures (social distancing, masking, handwashing etc) while allowing immunity to develop after being vaccinated. It is also still unclear what happens if an infected person is vaccinated, or in people who have no strong immune system (immunocompromised patients). “I have been vaccinated, can I now go, hugging and kissing on the street?” Sorry, you can’t. This is because you can still contract the virus even after being vaccinated: the body needs time to develop immunity against an infection. This is the main reason for jab-spacing as it enables the body to build an immune response.
Conclusion
Vaccines trigger the production of memory cells (T-lymphocytes and B-lymphocytes) to fight infections and can protect us from life-threatening diseases. The more people get vaccinated, the more likely we are to achieve herd-immunity, where even unvaccinated people are protected. A good example of herd-immunity is a burning bush. The fire keeps spreading but when it encounters a large space or a river, it stops spreading and the other side of the bush will not be affected. However, the space must be large enough to make this happen. Vaccinated individuals are like the large space or river, they help in stopping the spread of the disease to the unvaccinated hence, protecting the unvaccinated. Also, remember that for this kind of protection to occur, a sufficient number of individuals must have been vaccinated. There is no scientific basis to show that an mRNA-based vaccine can change your DNA. Developing an mRNA vaccine within a short period is a phenomenal advancement in science. As people are being vaccinated, every other protective measure is encouraged if we want to end this pandemic. We can only think of relaxing measures when the number of cases has considerably reduced. Stay safe and healthy, and together we are protected.
Acknowledgement
I would like to thank Dr Andrea Thorn, Dr Dale Tronrud, Dr Sam Horrell and Dr Yunyun Gao for their wonderful and helpful suggestions. Also, my thanks go to Dr Thomas Splettstoesser for making an image used for this post.
References
1. Regulatory Affairs Professionals Society. COVID-19 vaccine tracker [Internet]. [cited 2021 Jan 10]. Available from: https://www.raps.org/news-and-articles/news-articles/2020/3/covid-19-vaccine-tracker
2. Nature Communications. Vaccines work. Nat Commun [Internet]. 2018 Apr 24 [cited 2021 Jan 3];9. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5915378/
3. FDA. COVID-19 Vaccines. FDA [Internet]. 2021 Feb 18 [cited 2021 Feb 19]; Available from: https://www.fda.gov/emergency-preparedness-and-response/coronavirus-disease-2019-covid-19/covid-19-vaccines
4. Mueller B. U.K. Approves Pfizer Coronavirus Vaccine, a First in the West. The New York Times [Internet]. 2020 Dec 2 [cited 2021 Feb 19]; Available from: https://www.nytimes.com/2020/12/02/world/europe/pfizer-coronavirus-vaccine-approved-uk.html
5. World Health Organization. Vaccines and immunization [Internet]. [cited 2021 Jan 10]. Available from: https://www.who.int/health-topics/vaccines-and-immunization#tab=tab_1
6. Thèves C, Biagini P, Crubézy E. The rediscovery of smallpox. Clinical Microbiology and Infection. 2014 Mar;20(3):210–8.
7. Vitetta ES, Berton MT, Burger C, Kepron M, Lee WT, Yin XM. Memory B and T Cells. Annual Review of Immunology. 1991;9(1):193–217.
8. Centers for Disease Control and Prevention. Basics of Vaccines | CDC [Internet]. 2019 [cited 2021 Jan 10]. Available from: https://www.cdc.gov/vaccines/vpd/vpd-vac-basics.html
9. Sell S. How vaccines work: immune effector mechanisms and designer vaccines. Expert Rev Vaccines. 2019 Oct;18(10):993–1015.
10. Pardi N, Hogan MJ, Porter FW, Weissman D. mRNA vaccines — a new era in vaccinology. Nature Reviews Drug Discovery. 2018 Apr 1;17(4):261–79.
11. Zhang C, Maruggi G, Shan H, Li J. Advances in mRNA Vaccines for Infectious Diseases. Front Immunol [Internet]. 2019 Mar 27 [cited 2021 Jan 11];10. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6446947/
12. Jackson NAC, Kester KE, Casimiro D, Gurunathan S, DeRosa F. The promise of mRNA vaccines: a biotech and industrial perspective. npj Vaccines. 2020 Feb 4;5(1):11.
13. Martin C, Lowery D. mRNA vaccines: intellectual property landscape. Nat Rev Drug Discov. 2020 Sep;19(9):578–578.
14. Baden LR, Sahly HME, Essink B, Kotloff K, Frey S, Novak R, et al. Efficacy and Safety of the mRNA-1273 SARS-CoV-2 Vaccine. New England Journal of Medicine [Internet]. 2020 Dec 30 [cited 2021 Jan 10]; Available from: https://www.nejm.org/doi/10.1056/NEJMoa2035389
15. Sandbrink JB, Shattock RJ. RNA Vaccines: A Suitable Platform for Tackling Emerging Pandemics? Front Immunol. 2020;11:608460.
16. Pan American Health Organization WHO. COVID-19 Vaccine Explainer: COMIRNATY®, COVID-19 mRNA vaccine - PAHO/WHO | Pan American Health Organization [Internet]. [cited 2021 Feb 5]. Available from: https://www.paho.org/en/documents/covid-19-vaccine-explainer-comirnatyr-covid-19-mrna-vaccine

Was ist mit den Cap-Strukturen 5'UTR und 3'UTR des mRNA-Impfstoffes, welches nur das Struktur Protein codiert ? Sollen die fremden Viren mRNA-Stränge des Impfstoffes von den RNA-Sensoren (RIG-I, Mda-5, IFIT) erkannt werden ? Wird das translatierte Spike (S) Gen zu einem Protein auch an die MHC-I Moleküle gebunden, um es als Antigene zu präsentieren ?