Introduction
The short answer to this question is “almost certainly not”. However, we live in an unprecedented time; where people are both tired of experts while simultaneously believing that having read a meme on social media makes one an expert. So, what do I even mean by “almost certainly”? Between the politicians and the scientists on TV you’re probably tired of not getting a straight answer. I can’t speak for the politicians, but there is a reason for this from scientists. Scientists don’t like to work in absolutes. Not because we want to hide something, but because uncertainty is our home ground. Science works by minimising our uncertainty to the point where we can identify the simplest [1] and most likely outcome based on our observations.
So, was SARS-CoV-2 made in a lab? You can’t help but think: “It could have happened though, right?” This guy, Professor Nikolai Petrovsky from Flinders University, certainly thinks it’s possible. He also said it could be “…a chance transmission of a virus from an as yet unidentified animal to human”, but that’s not as interesting a headline. You can watch his full interview on the topic here.
[1] This is called the law of parsimony, or Occam’s razor, which states that the simplest solution is most likely the right one.
What will be discussed here?
Before I get into the science, I want to clarify the sort of claims I am addressing. As scientist we can only address a claim where the data are available and verifiable. Many of the arguments for the virus being created in the lab start with something along the lines of President Trump’s statement on April 30th.
“We have people looking at it very, very strongly. Scientific people, intelligence people, and others. We’re going to put it all together. I think we will have a very good answer eventually. And China might even tell us.”
President Donald Trump, April 2020
Ominous, but obviously lacking any real data. When pressed for evidence to prove his claims he retorted with
“I can’t tell you that. I’m not allowed to tell you that.”
President Donald Trump, April 2020
State secrets aside, this does not cut it in this battle ground. It is impossible to make a valid argument from secret data. This would be like me submitting a paper to a scientific journal and replying to reviewer’s comments with
“Just trust me, I have the data to back up my claim that alpacas breath fire when we’re not looking, but I’m not allowed to show you it because big wool is stopping me”
Dr Sam Horrell, June 2020.
Hence, we will only deal with claims for which there are valid data (See Figure 2).

The other argument typically sounds something like
“Of course it looks like the virus evolved naturally, these are very clever people that know how to cover their tracks”
Karen on Facebook, 2020.
This puts you in the unfortunate position of trying to prove a negative with your counter argument, which is, sadly, not possible. If we’re arguing this in a scientific manner, we must adhere to the burden of proof and provide positive evidence which allows us to find the simplest and most likely conclusion from our observations. A classic fallacy along the “very clever people” line is the creationist argument for a young Earth that God has made look old to trick the non-believers. Any science you try and throw at this is credited to God and a lack of faith, so you can’t argue this logically, but it does run into the problem of infinitely increasing complexity. You can see how were rocketing away from the simplest and most likely answer here.
On Natural Selection
Evolution and natural selection are central to this discussion. If you are of the opinion that evolution does not exist, then the rest of this article is not going to convince you and I hope you enjoyed the fire breathing alpaca picture. Natural selection works like this: each time a species reproduces there is a chance a mutation will occur in their genome. If this change grants an advantage (i.e. long necked giraffes), this increases the rate of survival and the chance that the giraffe has offspring allowing the change to persist in the population. If a change results in a considerable disadvantage (i.e. stumpy necked giraffes) it is less likely to be passed on to the next generation and will be selected out. Then there are some mutations which are innocuous and will persist in the genome. Although they are not useful to the species, they are very useful to evolutionary biologists when tracing a species’ genetic lineage. Viruses and bacteria have a considerable advantage when it comes to natural selection, as they reproduce at a much faster rate than us mammals. For example, E. coli cells can divide every 30 minutes, so will go through several generations over the course of a single day, which means a greater chance of stumbling onto a favourable mutation! Ever wonder why antibiotic resistance is such a big problem? Because of speedy evolution.
The Coronavirus Origin Story
The new Coronavirus SARS-CoV-2 was first identified after a pneumonia outbreak on the 12th of December 2019. Its genome was sequenced, and it showed 79.6% sequence identity to the virus causing Severe Acute Respiratory Syndrome (SARS) from 2002 - and 96% sequence identity to a bat coronavirus (RaTG13-CoV) which was recently reported by a lab in Wuhan. Since then all manner of conspiracy theories have popped up suggesting that this Coronavirus was produced in a lab in Wuhan, was intentionally or accidentally released, and had been specifically designed to target humans. And why not? 96% sounds too high to be a coincidence, right? Releasing this bat virus must be the cause of COVID-19! However, if we compare humans to one of their closest relatives, the chimpanzee, we can see that we also share 96% of our genomes. And as you can see from Figure three there are a fair few differences between us. Bringing it back to coronaviruses, that 96% difference accounts for 1,100 differences between these viruses. If we line up the sequences, we see a random distribution of mutations across the genome which follows the natural evolution typical of coronaviruses. We also have the benefit of previous data from the SARS-CoV outbreak in 2002. Human SARS-CoV was found to share 99.8% sequence identity with a palm civet coronavirus, with only 202 differences between the viruses. If this is the level of similarity that has been observed historically, it follows that a 96% identical virus is not likely to be the immediate source of a species jumping global pandemic. Even if it was the immediate source this only proves the virus has come from a bat, a species not known for their molecular biology expertise.
Figure 3: An accurate comparison of 96% identical species, Homo sapiens (left) and Pan troglodytes (right). Picture by Thomas Splettstößer.
Super Villain Interlude
If I was a super villain that had released bat corona virus aiming to shut down the world with a pandemic, I’d effectively be spinning an evolutionary roulette wheel and hoping it landed on unprecedented global health crisis. Not so much maniacal as just lucky. So, it’s highly unlikely (there’s that word again) that SARS-CoV-2 came directly from the bat coronavirus being released from the lab in Wuhan. If we stop for a moment and think about it, the bat corona virus already existed in the world, so what would releasing it from a lab without extensive modification really achieve? It is much more likely that there is an animal intermediate we’re currently missing in the natural evolution of Coronavirus, most likely the result of having animals in close proximity to other animals as well as humans at the animal market in Wuhan. But as of the writing of this blog this route has not been proven.
Still not convinced that the virus did not come from a lab? OK, let’s keep going. How do we even go about making a virus? At this point we are going to have to dig into some molecular biology, so hold on to your butts!
Homemade Viruses
We start with everyone’s favourite helical molecule, DNA, and a process called transcription. In transcription, DNA is partially unwound and a single stranded complementary (opposite) copy of the DNA sequence is produced, which we call RNA. RNA then is translated into proteins. When a virus infects a cell, it releases its genetic material (DNA or RNA) and uses our own cellular machinery to produce more viruses. If we were so inclined *cough super villain cough*, we could isolate this genetic material and, using an enzyme called reverse transcriptase, make a copy of the viral genome for our own nefarious purposes (or try and make a vaccine). This is called complimentary DNA (cDNA) and can be used to produce an infectious virus in a host which we can manipulate according to our wishes. In fact, this technique has been used already to study caliciviruses, alphaviruses, flaviviruses, arteriviruses, and *drum roll* coronaviruses! This paragraph makes this sound easy but don’t be fooled, this is certainly not the case. Making a zoonotic virus, an animal virus that can infect humans, is a significant undertaking, but not as significant as making a zoonotic virus that can be spread between humans.
So how do we know this is not where our SARS-CoV-2 comes from? To start, we are going to investigate the genome of SARS-CoV-2 and compare it with other notable coronaviruses. A recent paper published in Nature by Andersen and colleagues has identified two notable features in SARS-CoV-2’s genome that can help us answer this question. The first is that SARS-CoV-2 interacts well with a human protein called ACE2 because of five mutations on the spike protein (the bits poking out of the virus in Figure 4 – for more information on the spike protein see here). The second is that SARS-CoV-2’s spike protein has an additional twelve bases in its RNA sequence which make it particularly infectious and able to jump between host species. On face value, this sounds like a convincing argument for SARS-CoV-2 being made in a lab. Just add a little change to the genome and release it on an unsuspecting populace. Basic super villain stuff. However, as we dig a little deeper into the science behind this, this begins to seem much less likely.
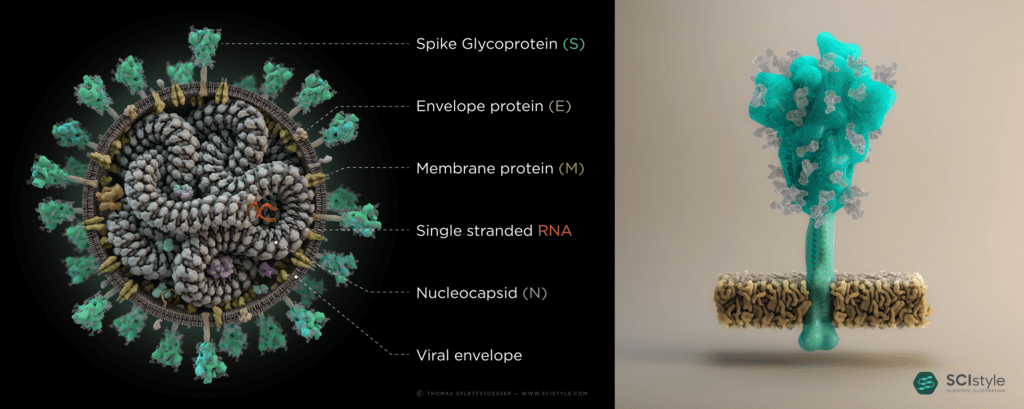
Figure 4: Illustration of SARS-CoV-2 and its spike protein by Thomas Splettstößer.
SARS-CoV-2 and ACE2 Binding
Let’s start with the optimised binding to human Angiotensin-Converting Enzyme 2, or ACE2 for short. ACE2 is a human enzyme that decorates the outer surface of a variety of cells throughout the human body, including the lungs. On a normal day, ACE2 plays an important role in cardiovascular (heart) and renal (kidney) function by producing vasodilators, key molecules that open blood vessels to increase blood flow and lower blood pressure. On an abnormal day an invasive virus (SARS-CoV-2) can bind to ACE2, enter our cells, and hijack our cell’s machinery to produce more viruses. If we compare the receptor binding domains of the spike protein from SARS-CoV (SARS-CoV-2’s 2002 predecessor), bat coronavirus and the SARS-CoV-2, we can see five key differences which improve SARS-CoV-2’s interaction with human ACE2. However, computational simulations show the interaction is far from perfect, and the binding differs from previously predicted binding modes. Furthermore, computational modelling suggests the spike protein is capable of recognising ACE2 in a number of animal species, with the exception of mice or rats. If these five key mutations were the only differences it would be more indicative of deliberate manipulation, however, the presence of 1095 other mutations distributed across the genome is much more suggestive of evolution through an animal intermediate.
Super Villain Interlude II: Electric Boogaloo
If I don my super villain costume again, to cover my tracks and make this look convincing I need to identify and isolate the bat corona virus, produce cDNA from that virus, develop a system to produce and study my new virus in a lab separate from current published methods, perform extensive computational modelling to identify a previously unreported binding mode for the spike protein, and then add in thousands of innocuous mutations without impairing the virus. Is all this possible? Of course, we have the technology as I explained earlier. But is it likely? Not really. This would take a large team of world leading experts from several different fields working for years in complete secrecy at the cutting edge of molecular biology. At this point were entering that rocky ground from earlier where the justification for the conspiracy theory is getting complex to the point of near impossibility.
Adding Sugar to a Virus Makes it Worse?
Next up, a polybasic furin cleavage site and O-linked glycans! Or, in English, some other stuff that makes SARS-CoV-2 more infectious. Part of SARS-CoV-2’s spike protein has a sequence made up of two different amino acids (RRAR) which is recognised and cut by a protease (a protein cutting enzyme) called furin. Cutting this sequence is predicted to be a key factor in virus binding to and gaining entry to cells. These sites are a signature of other highly infectious avian influenza viruses; affecting the pathogenicity of the virus and the hosts the viruses can infect. Natural selection of these sites can allow it to jump between species and turn a low-level pathogen to a highly pathogenic, ‘we-should-all-be-worried, “it’s over 9000”’-level pathogen.
What does that have to do with glycans? When furin cleaves the spike protein it makes two new sites either side of the cut, which scientist have predicted to be targets for O-linked glycosylation (attachment of a type of sugar to oxygen atoms on a protein). But what do these glycans even do? Well, we don’t exactly know yet for SARS-CoV-2. But we do know from experience that O-linked glycosylation can be used by viruses to avoid the immune system.
So, what does this cleavage site tell us about the possibility of making corona virus in a lab? The development of the furin cleavage site and the prediction of glycans also help us put this conspiracy theory to rest. Such cleavage sites are typically the result of a low-pathogenicity virus interacting with an immune system over many generations. Of course, we have the technology to add in the RRAR sequence into our hypothetical cDNA virus genome no problem, but accurately predicting where to put that site is a wholly different challenge. Natural selection in viruses can manage this by rolling the dice many millions of times until a random change, or more likely changes, grant such a significant advantage that a dominant version of the virus is selected out; a process that has been observed previously with influenza and furin cleavage sites. If you want a cleavage site for your new lab made virus, your best bet is to isolate a genetically similar virus and expose it repeatedly animals with ACE2 receptors akin to human ACE2. Cell culture wouldn’t cut it as interaction with an immune system is the driving factor in these changes, and we’ve already seen that rats and mice aren’t a viable system from the computational modelling. A piece of work on this scale represents a considerable time sink and monetary investment in an inefficient process which relies on roll of the dice to provide the desired results. As we have observed this evolutionary behaviour before in nature it stands to reason that the furin cleavage site is the result of natural selection and not deliberate manipulation.
Conclusion
We’ve covered a lot of ground from abductive reasoning and a young Earth to molecular biology and furin cleavage sites in our quest to unpick this conspiracy theory. As more studies are published the specifics of this may change, but, barring a colossal government coverup being unmasked, the involvement of deliberate manipulation in a lab appears unlikely. The evidence suggests the virus originated in bats, but it is highly unlikely the bat virus (RaTG13-CoV) is the direct precursor to SARS-CoV-2. Our best candidate for an intermediate species comes from a pangolin coronavirus which has been found to share the five mutations in the spike protein that facilitate ACE2 binding, but not the furin cleavage site11. We have shown that it is indeed possible to make our own viruses in a lab, but SARS-CoV-2’s backbone doesn’t match up with any of the currently available reverse genetic systems so this is unlikely to be a factor. And finally, looking deeper into the genome of SARS-CoV-2 we see ample evidence of natural selection across the whole viral genome, not just in the spike protein’s binding region, and the appearance of a furin cleavage site; a well-documented naturally selected phenomenon observed in viruses previously. Based on the available evidence, discounting any secret data that may be being held hostage in a secret lair hidden in a volcano, we come to the most logical and simple answer. SARS-CoV-2 was most likely not made in a lab but evolved naturally from bat coronavirus via an animal intermediate, possibly pangolins.
Further Reading
- The Proximal origin of SARS-CoV2
- A genome-wide comparison of recent chimpanzee and human segmental duplications
- No credible evidence supporting claims of the laboratory engineering of SARS-CoV-2
- Bacterial artificial chromosomes: A functional genomics tool for the study of positive-strand RNA viruses
- Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein
- Receptor recognition by the novel coronavirus from Wuhan: An analysis based on decade-long structural studies of SARS coronavirus
- Coronavirus reverse genetic systems: Infectious clones and replicons
- Trypsin treatment unlocks barrier for zoonotic bat coronavirus infection
- History of highly pathogenic avian influenza
- Global aspects of viral glycosylation
- Probable pangolin origin of SARS-CoV-2 associated with the COVID-19 outbreak
Acknowledgements
I would like to thank a number of people for help with the writing of this post, Harri Webb for acting as a fire breathing alpaca wrangler, Mary Cruise for proof reading and suggestions, Thomas Splettstöße for the figures that look professionally made, and the members of the Coronavirus structural taskforce, particularly Alex Payne, Dale Tronrud, and Andrea Thorn for all their help and suggestions.

What a wonderful article, dear Sam!
I really loved to read it. Especially the "God wanted the earth to look old" argument and your super villian point of view, mixed up with lots of microbilogical scientific stuff – well, ok, i didn' get it all... but the fact that i even tried to, was only because of your great style of writing 😀
Thanks for this great reading time.
Holger.
Thanks Holger, I'm really glad you enjoyed reading it.
Hello, I have read parts of your excellent article and will study it a few more times. It seems to be fairly common for virology labs to study in vitro evolution/selection of viruses by multiple-passaging, etc, and it is not unheard of for the beasties to escape, indeed it happened just down the road from where I am now (;-):
https://www.newscientist.com/article/dn12615-faulty-pipe-blamed-for-uk-foot-and-mouth-outbreak/
Anyway, keep up the great work.
P.S. As a favour could you possibly explain this sentence a bit more: "This puts you in the unfortunate position of trying to prove a negative with your counter argument, which is, sadly, not possible."
Hi Jon,
Thanks very much for the question. Yes a virology lab will need to produce viruses to be able to study them, and it is certainly possible for a virology lab to make a mistake and for a virus to be released. Depending on how dangerous the virus you are working on is there will be levels of safety measures to keep it all in check, but humans (even scientists) make mistakes. The main point I would like to get across with this article is that all the evidence points to the natural evolution of the virus, the virus was already in the environment in bats and natural mutation has allowed it to make the species jump to humans. This is not to say that a lab couldn't have produced it and accidently released it, it just seems very unlikely given the evidence I outlined in the article.
As for the proving a negative, I've take this from a Wikipedia article about proving a negative. "The difference with a positive claim is that it takes only a single example to demonstrate such a positive assertion ("there is a chair in this room," requires pointing to a single chair), while the inability to give examples demonstrates that the speaker has not yet found or noticed examples rather than demonstrates that no examples exist (the negative claim that a species is extinct may be disproved by a single surviving example or proven with omniscience)."
You could also look up arguing from ignorance if you're interested.
Thank you, only just saw your reply. Re: "This puts you in the unfortunate position of trying to prove a negative..." so the negative is that the virus did not evolve naturally, or virus ¬ evolved naturally, right ;-? I'm rusty on truth tables ;-0 Re: "... trying to prove a negative with your counter argument, which is... not possible." Ok, so do you mean that you cannot prove virus ¬ evolved naturally by "a lack of contrary evidence" (wikipedia article "Argument from ignorance") i.e. you mean you can't prove virus ¬ evolved naturally by a lack of evidence that it did evolve naturally?? No worries, I think that's what you mean. Ignoring all the bioinformatics, etc, I guess to prove it could have evolved naturally you would have to take a virus from a bat and show it infects people? Maybe that has been done?
So the proving the negative comes into play when someone replies to your evidence countering their argument with something like the text quoted above ("Of course it looks like the virus evolved naturally, these are very clever people that know how to cover their tracks"). If they are making this argument you are then forced to prove they are not covering their tracks (proving the negative) when they should actually be providing evidence for the fraud if this is the case. It's not a valid argument to just say someone is so clever that they have everyone fooled, it's impossible to prove anything when evidence you present is rebutted with "that's just been cleverly done to trick you".
The bioinformatics shows that the current virus that infects humans likely started with the bat variant, a few mutations down the line we have a species jump and an increase in virulence which ended us up in the current pandemic. The original bat virus couldn't infect humans and had likely been kicking about in the natural world for a long time. The mutations are the next stage in its evolution, like the primates that became man, both are still around in some form but they have now diverged.
What a great article! Thank you!
So, it is unlikely that SARS-COV-2 is an artificial lab product.
However, don´t you underestimate the capacity of the super villain a bit, especially if it´s a government with nearly unlimitied funding? It has been done before. By chance i suddenly came across this:
https://media.defense.gov/2019/Apr/11/2002115480/-1/-1/0/14NEXTGENBIOWEAPONS.PDF
and it made my stomach revolt.