In addition to mRNA vaccines, another type of vaccine is employed against COVID-19: Vector vaccines—like the one from AstraZeneca—contain a mostly functioning virus. But how do they work exactly? What are their strengths and weaknesses? And finally, are they safe?
The novel mRNA vaccines against SARS-CoV-2 have caused some controversy, and many concerns have been raised. According to these, the vaccine has not been sufficiently tested, long-term effects are yet unknown, the vaccine might affect the patient’s genetic material.
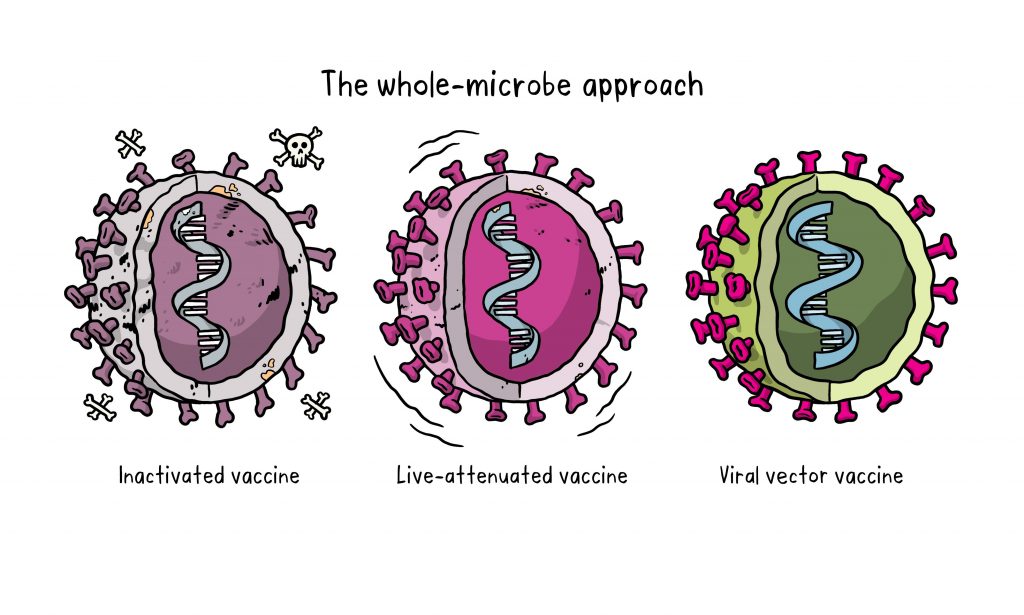
While mRNA vaccines enfold the genetic material in a tiny lipid particle, so-called viral vector vaccines put it in a viral shell that functions as a transporter (vector). A large number of vaccines of this type are under development, and they are all based on various strains of the same virus known for causing the common cold. The European Medicines Agency (EMA) recommended the Oxford–AstraZeneca vaccine for approval in late January, and the United States approved the use of a similar vaccine from Johnson & Johnson in February. Why?
Viruses are transporters for genes
Viruses are not considered living organisms because they have no metabolism of their own. They contain only the genes that cause the host cell to produce new viruses.
In the evolution of viruses, some of their skills have become exceptionally refined. Viruses can specifically infect host cells, escape the host’s immune defenses, and insert their genetic material into the host cell. Once molecular biologists recognized these capabilities, scientists could turn viruses into tools that do not cause disease through genetic engineering. As modified tools, they are referred to as viral vectors.
Viruses as tools
To make viruses useful for research purposes, they are modified. Their natural genes must be changed or deleted. What is left in the end is a transporter that can be used to introduce desired genes into a host. This method is used, for example, to produce transgenic (genetically modified) plants or cell lines. In this process, researchers usually keep some natural functions of the virus such as the capability to invade host cells.
Viruses as vaccines
What exactly must be changed to turn a virus into a vaccine?
In order not to cause a disease, the virus must lose its ability to multiply inside the body. Viruses without the ability to reproduce are called "replication-deficient". In the past, replication was disabled via lengthy cell culture techniques but, since the 2000s, genetic engineering has made it possible to selectively remove or modify genes of a virus to stop it from reproducing. Removing critical genes can completely prevent reproduction of the virus and any chance of reversion to the pathogenic (disease-causing) variant can be prevented [3].
Gene modification can also be used to introduce structures such as the spike gene of SARS-CoV-2 into a virus that is harmless to humans. This addition turns the virus into a vaccine.
Viruses have been used as tools for a long time. Viral vectors were first described in 1972 [1]. In the early 1990s, gene transfer was first used for therapy through an attenuated adenovirus as vector [2].
Across the last twenty years, a number of vaccines based on viruses have been developed. An example of an approved vector vaccine is the Ebola vaccine Ervebo [4]. Because there is no treatment for Ebola to date, the epidemics from 2013 to 2020 were devastating for Central Africa. Mortality rates ranged from 25 to 90% [5]. In August 2018, Ervebo was used for the first time to vaccinate a large group of people. It proved to be highly effective [6] and was also licensed for employment in the EU in 2019 [5].
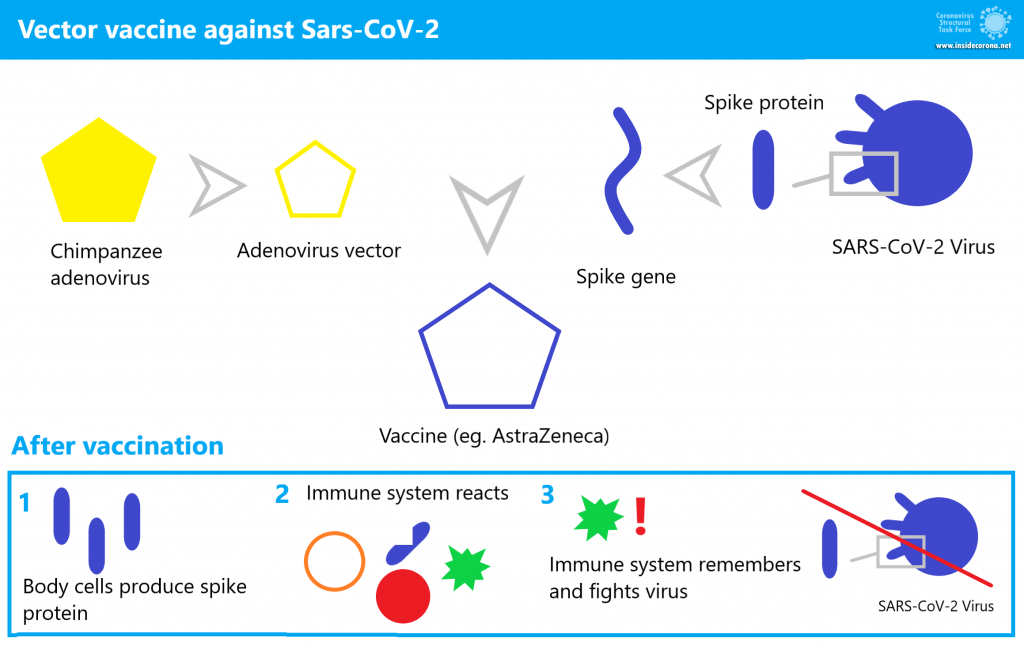
How does the AstraZeneca vaccine work?
This vaccine is known as AZD1222 (also called ChAdOx1 nCoV-19) [8]. It is an adenovirus that is similar to a pathogen for the common cold, but this strain was originally found in chimpanzees. It has been modified to be replication-deficient. AZD1222 was also modified to contain the coronavirus’ surface protein (spike) gene [9]. This chimpanzee virus is used because most people’s immune systems are familiar with adenoviruses that infect humans, and their immune systems might respond to those viruses before they can infect cells and produce spike protein. In comparison, the chimpanzee virus is unfamilar to the human immune system and is hence capable of infecting a cell [10]. Side effects can occur, but they do not stem from the actual disease. Feeling unwell after being vaccinated is triggered by our immune system fighting against an intruder.
AZD1222 invades some human host cells just like a usual common cold virus. During this process, the vector transports the gene of the spike protein into the host cell. The spike DNA is utilized by the cell to produce spike protein. This spike protein acts as an antigen, a substance that triggers an immune response. The immune system recognizes it as foreign and starts attacking, antibodies are produced and the few "infected" cells containing AZD1222 genetic material are destroyed. Antibodies bind to the antigen like pieces of a puzzle and hold on to it. The antigen-antibody clumps are then degraded.
What remains from the whole process are memory cells that recognize the spike protein when attacked by the real SARS-CoV-2. Previous training with the vaccine makes the next immune response faster and more effective. This prevents disease before it can break out.
How effective and safe is the vaccine?
AZD1222 has already been tested in over 20,000 people across three phases of clinical trial [11]. Since its approval, data has been continued to be collected and combined into additional studies. Through these, it is possible to investigate possible side effects even more thoroughly since millions of people are being vaccinated and observed in the "real world".
One example is a British study on efficacy in people over 70 years of age. The result: A single dose of AstreZeneca’s viral vector vaccine or BioNTech’s mRNA-based vaccine is effective against a symptomatic Corona infection in 60-75% of cases. The likelihood of hospitalization is reduced by 80% with either vaccine [12].
The first nationwide study took place in Scotland. It was led by the University of Edinburgh and collected data from 5.4 million vaccinated people. Preliminary data show that AstraZeneca’s single-dose vaccine prevented a severe course—requiring hospitalization—in up to 94% of cases [13].
Through these real world studies, it is now known that the vector vaccine shows greater effectiveness with a larger interval between the first and second dose. Therefore, these studies recommend an interval of twelve weeks [14].
In terms of side effects, some data has already been collected on AZD1222 in the UK due to the earlier start of vaccination [15]. Common side effects experienced by one in ten patients receiving vaccination include:
- tenderness, pain, warmth, itching, or bruising at the injection site,
- feeling generally unwell or tired,
- chills or feverish feeling,
- headache,
- nausea, and
- joint pain or muscle soreness.
These frequent side effects are similar to those of the BioNtech vaccine. However, no serious side effects have been detected in the trials. Fever or flu-like symptoms are less common and are a general sign of the vaccine activating the immune system and therefore taking effect. In any case, these symptoms are not comparable to the risks of a severe COVID-19 course.
The EMA is currently looking into cases of blood clotting that occurred shortly after an AstraZeneca vaccination. So far there are no indications that the vaccine is unsafe. The frequency of blood clots in vaccinated people is not higher than it generally is in the population. According to AstraZeneca [15.1] the risk of pulmonary embolism, deep vein thrombosis (DVT) or thrombocytopenia is not higher in people receiving the vaccine compared to the general public. Among 17 million vaccinated patients in Europe, some are expected to suffer from these diseases, but it is very likely not linked to the vaccine.
What is in the vaccine?
In addition to the modified adenovirus, the vaccine contains several other substances [16].
Most are additives that stabilize the vaccine and make administration easier. These include amino acids, stabilizers, alcohol, sugar, salt, binders, and water. In addition, the vaccine is free of food allergens (such as soy or lactose) and contains no ingredients of human or animal origin. That sounds incredible for a chimpanzee virus that needs a host to reproduce. How can this be?
All modifications and amplification of the virus took place in a human cell line used in the laboratory for such purposes, so-called HEK293 cells. These cells are, however, not part of the vaccine.
What are the advantages of this type of vaccine?
Viral vector vaccines—just plain RNA/DNA vaccines—can be developed very quickly. The reason: As soon as the genes of the pathogen are known, they can be used for vaccine development. This shortens the time until clinical trials can begin. This makes vector vaccines suitable for sudden epidemic outbreaks [18].
One advantage of the AstraZeneca vaccine is its dosage. Oxford–AstraZeneca initially tested different dosing approaches for their vector vaccine. Vaccinations with two standard doses at intervals of four to twelve weeks and a vaccination with only one dose were compared. For this, a standard dose contains 5x1010 virus particles [19].
"By using a more effective dosing regimen," said Professor Pollard, the lead scientist at Oxford, "more people could be served with the same amount of vaccine."[20]. The ideal regimen was found to be the administration of a half dose (2.2x1010 virus particles) followed by a standard dose at least one month apart. The efficiency in this case was 90% [21]. However, this must first be confirmed by additional data. So far, vaccination is done with two standard doses.
Conclusion
In nature, viruses have perfected the ability to insert their genetic material into hosts. The fascinating abilities of viruses can nowadays be used by scientists. They are therefore used to create genetically modified plants or cells, to treat hereditary diseases, and to produce vaccines.
Vector vaccines—such as AstraZeneca’s—contain harmless and non-reproducing viruses that contain a part of a disease-causing virus. By artificially "infecting" the patient with the vector vaccine, the immune system is trained to respond to the pathogen and can react more quickly and effectively in the event of a real infection in the future.
[1] Jackson D.A., Symons R.H., Berg P. Biochemical method for inserting new genetic information into DNA of Simian Virus 40: Circular SV40 DNA molecules containing lambda phage genes and the galactose operon of Escherichia coli. Proc. Natl. Acad. Sci. USA. 1972;69:2904–2909. doi: 10.1073/pnas.69.10.2904.
[2] Zabner, Joseph et al. 1993 Adenovirus-mediated gene transfer transiently corrects the chloride transport defect in nasal epithelia of patients with cystic fibrosis Cell, Volume 75, Issue 2, 207 – 216.
[3] Using directed attenuation to enhance vaccine immunity, Rustom Antia, Hasan Ahmed, James J Bull, bioRxiv 2020.03.22.002188; doi: https://doi.org/10.1101/2020.03.22.002188
[4] https://www.who.int/vaccine_safety/committee/topics/ebola/Jul_2019/en/
[5] https://www.ema.europa.eu/en/news/first-vaccine-protect-against-ebola
[6] https://www.thelancet.com/journals/lancet/article/PIIS0140-6736(16)32621-6/fulltext#seccestitle160
[7] https://www.nature.com/articles/d42859-020-00016-5
[8] https://www.ema.europa.eu/en/documents/product-information/covid-19-vaccine-astrazeneca-product-information-approved-chmp-29-january-2021-pending-endorsement_en.pdf
[9] https://jvi.asm.org/content/jvi/77/16/8801.full.pdf
[10] https://www.cdc.gov/vaccines/covid-19/hcp/viral-vector-vaccine-basics.html
[11] https://clinicaltrials.gov/ct2/show/NCT04516746
[12] https://www.medrxiv.org/content/10.1101/2021.03.01.21252652v1
[13] https://www.ed.ac.uk/files/atoms/files/scotland_firstvaccinedata_preprint.pdf
[14] https://www.rki.de/DE/Content/Kommissionen/STIKO/Empfehlungen/AstraZeneca-Impfstoff.html
[15] https://www.gov.uk/government/publications/regulatory-approval-of-pfizer-biontech-vaccine-for-covid-19/information-for-uk-recipients-on-pfizerbiontech-covid-19-vaccine#side-effects
[15.1] https://www.astrazeneca.com/media-centre/press-releases/2021/update-on-the-safety-of-covid-19-vaccine-astrazeneca.html
[16] https://www.astrazeneca.at/content/dam/az-at/pdf/2021/Vaccine%20guide%20for%20HCPs%20-%202021-02-02.pdf
[17] Draper, S., Heeney, J. Viruses as vaccine vectors for infectious diseases and cancer. Nat Rev Microbiol 8, 62–73 (2010). https://doi.org/10.1038/nrmicro2240
[18] https://www.frontiersin.org/articles/10.3389/fimmu.2018.01963/full
[19] https://papers.ssrn.com/sol3/papers.cfm?abstract_id=3777268
[20] https://www.astrazeneca.com/media-centre/press-releases/2020/azd1222hlr.html
[21] https://www.thelancet.com/journals/lancet/article/PIIS0140-6736(20)32661-1/fulltext
